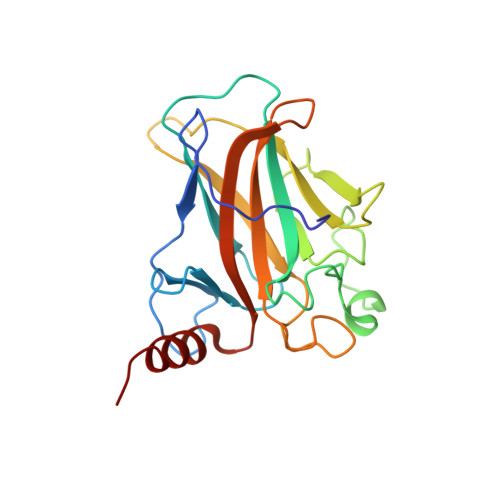
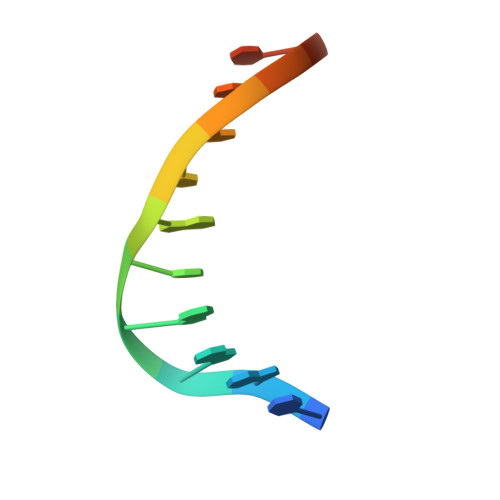
Structural Basis of DNA Recognition by p53 Tetramers
Kitayner, M., Rozenberg, H., Kessler, N., Rabinovich, D., Shaulov, L., Haran, T.E., Shakked, Z.(2006) Mol Cell 22: 741-753
- PubMed: 16793544 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2006.05.015
- Primary Citation Related Structures:
2AC0, 2ADY, 2AHI, 2ATA - PubMed Abstract:
The tumor-suppressor protein p53 is among the most effective of the cell's natural defenses against cancer. In response to cellular stress, p53 binds as a tetramer to diverse DNA targets containing two decameric half-sites, thereby activating the expression of genes involved in cell-cycle arrest or apoptosis. Here we present high-resolution crystal structures of sequence-specific complexes between the core domain of human p53 and different DNA half-sites. In all structures, four p53 molecules self-assemble on two DNA half-sites to form a tetramer that is a dimer of dimers, stabilized by protein-protein and base-stacking interactions. The protein-DNA interface varies as a function of the specific base sequence in correlation with the measured binding affinities of the complexes. The new data establish a structural framework for understanding the mechanisms of specificity, affinity, and cooperativity of DNA binding by p53 and suggest a model for its regulation by regions outside the sequence-specific DNA binding domain.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100.
Organizational Affiliation: