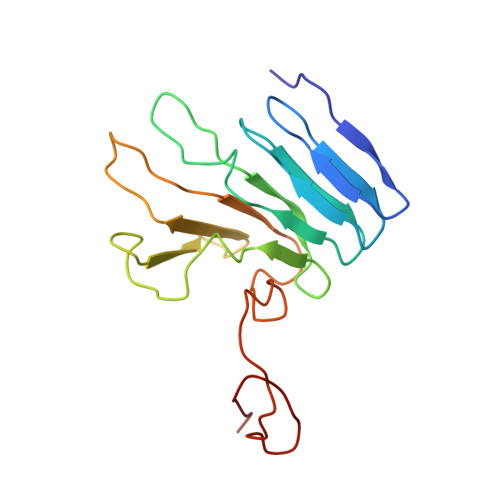
NMR structure of the R-module: a parallel beta-roll subunit from an Azotobacter vinelandii mannuronan C-5 epimerase.
Aachmann, F.L., Svanem, B.I., Guntert, P., Petersen, S.B., Valla, S., Wimmer, R.(2006) J Biological Chem 281: 7350-7356
- PubMed: 16407237 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M510069200
- Primary Citation Related Structures:
2AGM - PubMed Abstract:
In the bacterium Azotobacter vinelandii, a family of seven secreted and calcium-dependent mannuronan C-5 epimerases (AlgE1-7) has been identified. These epimerases are responsible for the epimerization of beta-d-mannuronic acid to alpha-l-guluronic acid in alginate polymers. The epimerases consist of two types of structural modules, designated A (one or two copies) and R (one to seven copies). The structure of the catalytically active A-module from the smallest epimerase AlgE4 (consisting of AR) has been solved recently. This paper describes the NMR structure of the R-module from AlgE4 and its titration with a substrate analogue and paramagnetic thulium ions. The R-module folds into a right-handed parallel beta-roll. The overall shape of the R-module is an elongated molecule with a positively charged patch that interacts with the substrate. Titration of the R-module with thulium indicated possible calcium binding sites in the loops formed by the nonarepeat sequences in the N-terminal part of the molecule and the importance of calcium binding for the stability of the R-module. Structure calculations showed that calcium ions can be incorporated in these loops without structural violations and changes. Based on the structure and the electrostatic surface potential of both the A- and R-module from AlgE4, a model for the appearance of the whole protein is proposed.
- Department of Life Sciences, Aalborg University, Sohngaardsholmsvej 49, DK-9000 Aalborg, Denmark.
Organizational Affiliation: