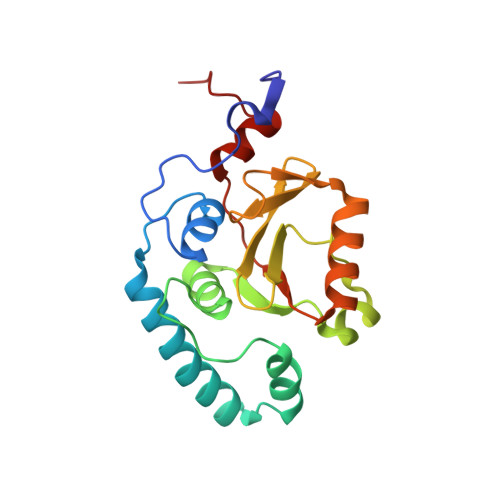
Deubiquitinating function of ataxin-3: insights from the solution structure of the Josephin domain.
Mao, Y., Senic-Matuglia, F., Di Fiore, P.P., Polo, S., Hodsdon, M.E., De Camilli, P.(2005) Proc Natl Acad Sci U S A 102: 12700-12705
- PubMed: 16118278 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0506344102
- Primary Citation Related Structures:
2AGA - PubMed Abstract:
Spinocerebellar ataxia type 3 is a human neurodegenerative disease resulting from polyglutamine tract expansion. The affected protein, ataxin-3, which contains an N-terminal Josephin domain followed by tandem ubiquitin (Ub)-interacting motifs (UIMs) and a polyglutamine stretch, has been implicated in the function of the Ub proteasome system. NMR-based structural analysis has now revealed that the Josephin domain binds Ub and has a papain-like fold that is reminiscent of that of other deubiquitinases, despite primary sequence divergence but consistent with its deubiqutinating activity. Mutation of the catalytic Cys enhances the stability of a complex between ataxin-3 and polyubiquitinated proteins. This effect depends on the integrity of the UIM region, suggesting that the UIMs are bound to the substrate polyubiquitin during catalysis. We propose that ataxin-3 functions as a polyubiquitin chain-editing enzyme.
- Howard Hughes Medical Institute and Department of Cell Biology, Boyer Center for Molecular Medicine, Yale University School of Medicine, New Haven, CT 06510, USA.
Organizational Affiliation: