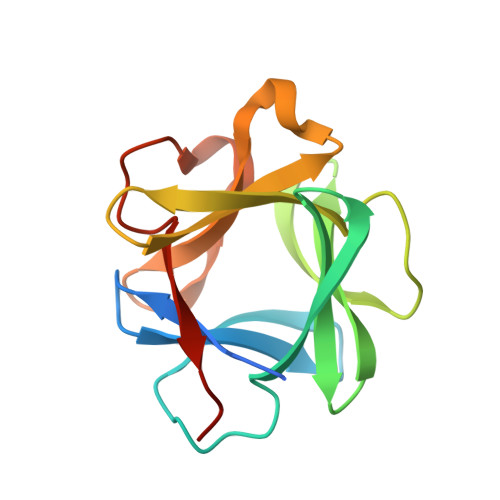
X-ray crystal structure of human acidic fibroblast growth factor.
Blaber, M., DiSalvo, J., Thomas, K.A.(1996) Biochemistry 35: 2086-2094
- PubMed: 8652550 Search on PubMed
- DOI: https://doi.org/10.1021/bi9521755
- Primary Citation Related Structures:
2AFG - PubMed Abstract:
Fibroblast growth factors (FGFs) are mitogenic and chemotactic agents for a wide variety of cell types and play a primary role in the regulation of angiogenesis. Angiogenesis is involved in a variety of critical physiological events including organogenesis, wound healing, ischemic collateral circulation, and solid tumor growth. High-resolution structural information is key to understanding the mechanism of action of these growth factors. We report here the X-ray crystal structure of human acidic FGF (aFGF), with data extending to 2.0 angstroms resolution. The crystal contains four independent molecules in the asymmetric unit. Each molecule contains a single bound sulfate ion, in similar juxtapositions. The bound sulfate is stabilized through hydrogen-bond interactions with residues Asn 18, Lys 113, and Lys 118 and defines a potential heparin binding site. The hydrogen bond with the N delta 2 moiety of Asn 18 appears to be the most conserved interaction, being similar to those observed for sulfate ion bound to human basic FGF (bFGF) and similar but not identical to interactions observed for bovine aFGF with heparin analogs. Of the added solvent groups, five ordered water molecules are conserved in each of the four independent structures of human aFGF. These water molecules, located at buried positions, provide hydrogen bonding partnerships with several buried polar groups in the core of the protein. A central interior cavity exists in each of the four structures, with sizes ranging from approximately 20 to 50 angstroms3. The cavity sizes appear to be significantly smaller than that observed in the related protein interleukin-1 beta. The region comprising the high affinity FGF receptor binding site is structurally very similar to the corresponding region from human bFGF, whereas the low affinity site is structurally quite different. The results provide a structural basis for the role of the low affinity binding site in FGF receptor discrimination.
- Institute of Molecular Biophysics and Department of Chemistry, Florida State University, Tallahassee 32306-3015, USA.
Organizational Affiliation: