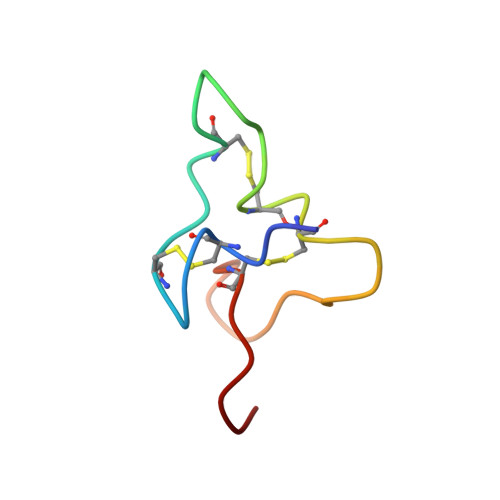
Structure of the fifth EGF-like domain of thrombomodulin: An EGF-like domain with a novel disulfide-bonding pattern.
Sampoli Benitez, B.A., Hunter, M.J., Meininger, D.P., Komives, E.A.(1997) J Mol Biology 273: 913-926
- PubMed: 9367781 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1356
- Primary Citation Related Structures:
1ADX, 2ADX - PubMed Abstract:
The structure of the fifth EGF-like domain (residues Q387 to E426) of thrombomodulin (TMEGF5) has been determined by two-dimensional NMR. TMEGF5 binds to thrombin with a Ki of 1.9 microM and has been shown to have a novel disulfide bonding pattern in a fully active fragment of TM. In EGF, the disulfide bonding pattern is (1-3,2-4, 5-6), while TMEGF5 has an uncrossed (1-2,3-4,5-6) pattern. The structure of this novel domain, determined from 483 NOE-derived distance restraints, appears to have diverged from the common EGF-like structure. Superposition of the 14 lowest-energy structures of TMEGF5 gives an overall r.m.s.d. of 1.09 A for the backbone atoms. The central two-stranded beta-sheet common to all EGF-like domains is not present in TMEGF5. The A loop, residues C390 to C395, is twisted away from interacting with the B loop, residues C399 to C407, as in EGF, and is close to the C loop, residues C409 to C421. This twist causes the N and C termini to be closer together in TMEGF5 than in EGF. Most of the residues that are important for activity lie on one face of the molecule, which is likely to be the thrombin-binding surface of the domain. The structure of the C loop within the domain, which is a beta-hairpin similar to EGF, is similar to the structure of a synthetic version of the loop bound to thrombin as determined by transferred NOE experiments. Despite the similarity in the structures of the loops, the residues immediately following C421 are in different positions in the two structures suggesting that these "tail" residues may change conformation upon thrombin binding.
- Department of Chemistry and Biochemistry, University of California, San Diego, CA 92093-0601, USA.
Organizational Affiliation: