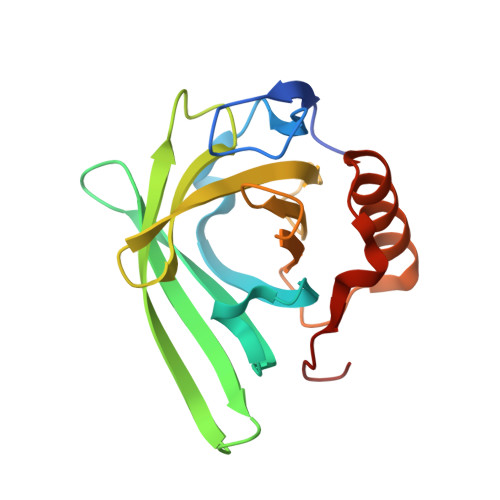
The membrane bound bacterial lipocalin Blc is a functional dimer with binding preference for lysophospholipids.
Campanacci, V., Bishop, R.E., Blangy, S., Tegoni, M., Cambillau, C.(2006) FEBS Lett 580: 4877-4883
- PubMed: 16920109 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.febslet.2006.07.086
- Primary Citation Related Structures:
2ACO - PubMed Abstract:
Lipocalins, a widespread multifunctional family of small proteins (15-25kDa) have been first described in eukaryotes and more recently in Gram-negative bacteria. Bacterial lipocalins belonging to class I are outer membrane lipoproteins, among which Blc from E. coli is the better studied. Blc is expressed under conditions of starvation and high osmolarity, conditions known to exert stress on the cell envelope. The structure of Blc that we have previously solved (V. Campanacci, D. Nurizzo, S. Spinelli, C. Valencia, M. Tegoni, C. Cambillau, FEBS Lett. 562 (2004) 183-188.) suggested its possible role in binding fatty acids or phospholipids. Both physiological and structural data on Blc, therefore, point to a role in storage or transport of lipids necessary for membrane maintenance. In order to further document this hypothesis for Blc function, we have performed binding studies using fluorescence quenching experiments. Our results indicate that dimeric Blc binds fatty acids and phospholipids in a micromolar K(d) range. The crystal structure of Blc with vaccenic acid, an unsaturated C18 fatty acid, reveals that the binding site spans across the Blc dimer, opposite to its membrane anchored face. An exposed unfilled pocket seemingly suited to bind a polar group attached to the fatty acid prompted us to investigate lyso-phospholipids, which were found to bind in a nanomolar K(d) range. We discuss these findings in terms of a potential role for Blc in the metabolism of lysophospholipids generated in the bacterial outer membrane.
- Architecture et Fonction des Macromolecules Biologiques, UMR 6098, CNRS-Universités Aix-Marseille I & II, Campus de Luminy, Case 932, 163 Avenue de Luminy, 13288 Marseille Cedex 09, France.
Organizational Affiliation: