Study of a major intermediate in the oxidative folding of leech carboxypeptidase inhibitor: contribution of the fourth disulfide bond
Arolas, J.L., Popowicz, G.M., Bronsoms, S., Aviles, F.X., Huber, R., Holak, T.A., Ventura, S.(2005) J Mol Biology 352: 961-975
- PubMed: 16126224 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.07.065
- Primary Citation Related Structures:
2ABZ - PubMed Abstract:
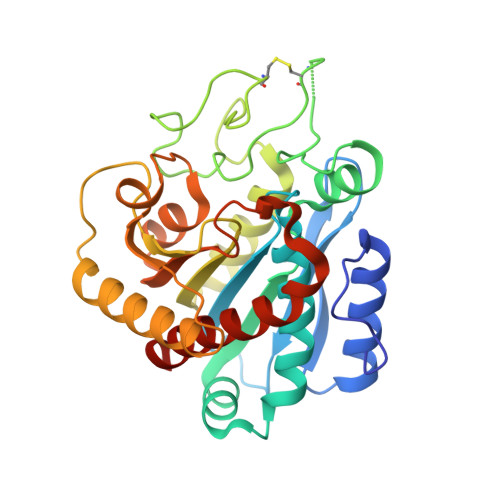
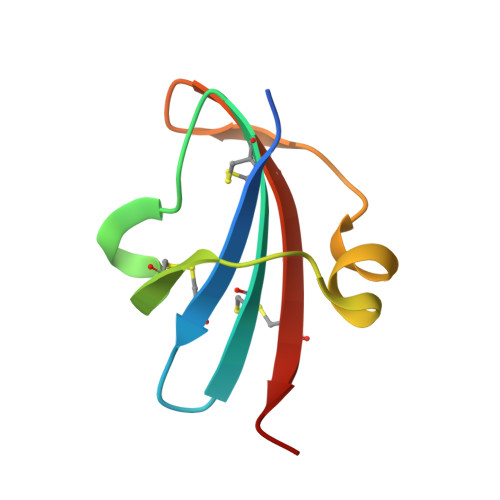
The oxidative folding pathway of leech carboxypeptidase inhibitor (LCI; four disulfide bonds) proceeds through the formation of two major intermediates (III-A and III-B) that contain three native disulfide bonds and act as strong kinetic traps in the folding process. The III-B intermediate lacks the Cys19-Cys43 disulfide bond that links the beta-sheet core with the alpha-helix in wild-type LCI. Here, an analog of this intermediate was constructed by replacing Cys19 and Cys43 with alanine residues. Its oxidative folding follows a rapid sequential flow through one, two, and three disulfide species to reach the native form; the low accumulation of two disulfide intermediates and three disulfide (scrambled) isomers accounts for a highly efficient reaction. The three-dimensional structure of this analog, alone and in complex with carboxypeptidase A (CPA), was determined by X-ray crystallography at 2.2A resolution. Its overall structure is very similar to that of wild-type LCI, although the residues in the region adjacent to the mutation sites show an increased flexibility, which is strongly reduced upon binding to CPA. The structure of the complex also demonstrates that the analog and the wild-type LCI bind to the enzyme in the same manner, as expected by their inhibitory capabilities, which were similar for all enzymes tested. Equilibrium unfolding experiments showed that this mutant is destabilized by approximately 1.5 kcal mol(-1) (40%) relative to the wild-type protein. Together, the data indicate that the fourth disulfide bond provides LCI with both high stability and structural specificity.
- Institut de Biotecnologia i de Biomedicina and Departament de Bioquímica i Biologia Molecular, Universitat Autònoma de Barcelona, 08193 Bellaterra, Spain.
Organizational Affiliation: