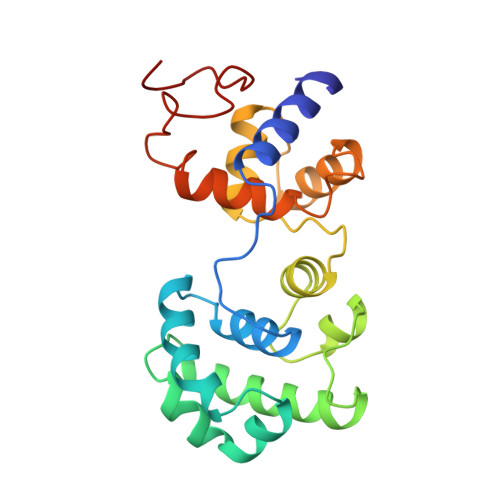
Novel DNA binding motifs in the DNA repair enzyme endonuclease III crystal structure.
Thayer, M.M., Ahern, H., Xing, D., Cunningham, R.P., Tainer, J.A.(1995) EMBO J 14: 4108-4120
- PubMed: 7664751 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1995.tb00083.x
- Primary Citation Related Structures:
2ABK - PubMed Abstract:
The 1.85 A crystal structure of endonuclease III, combined with mutational analysis, suggests the structural basis for the DNA binding and catalytic activity of the enzyme. Helix-hairpin-helix (HhH) and [4Fe-4S] cluster loop (FCL) motifs, which we have named for their secondary structure, bracket the cleft separating the two alpha-helical domains of the enzyme. These two novel DNA binding motifs and the solvent-filled pocket in the cleft between them all lie within a positively charged and sequence-conserved surface region. Lys120 and Asp138, both shown by mutagenesis to be catalytically important, lie at the mouth of this pocket, suggesting that this pocket is part of the active site. The positions of the HhH motif and protruding FCL motif, which contains the DNA binding residue Lys191, can accommodate B-form DNA, with a flipped-out base bound within the active site pocket. The identification of HhH and FCL sequence patterns in other DNA binding proteins suggests that these motifs may be a recurrent structural theme for DNA binding proteins.
- Scripps Research Institute, Department of Molecular Biology-MB4, La Jolla, CA 92037, USA.
Organizational Affiliation: