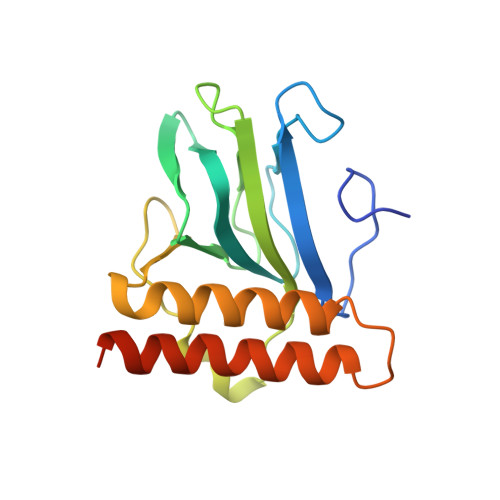
The 1.6 A crystal structure of the AraC sugar-binding and dimerization domain complexed with D-fucose.
Soisson, S.M., MacDougall-Shackleton, B., Schleif, R., Wolberger, C.(1997) J Mol Biology 273: 226-237
- PubMed: 9367758 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1314
- Primary Citation Related Structures:
2AAC - PubMed Abstract:
The crystal structure of the sugar-binding and dimerization domain of the Escherichia coli gene regulatory protein, AraC, has been determined in complex with the competitive inhibitor D-fucose at pH 5.5 to a resolution of 1.6 A. An in-depth analysis shows that the structural basis for AraC carbohydrate specificity arises from the precise arrangement of hydrogen bond-forming protein side-chains around the bound sugar molecule. van der Waals interactions also contribute to the epimeric and anomeric selectivity of the protein. The methyl group of D-fucose is accommodated by small side-chain movements in the sugar-binding site that result in a slight distortion in the positioning of the amino-terminal arm. A comparison of this structure with the 1.5 A structure of AraC complexed with L-arabinose at neutral pH surprisingly revealed very small structural changes between the two complexes. Based on solution data, we suspect that the low pH used to crystallize the fucose complex affected the structure, and speculate about the nature of the changes between pH 5.5 and neutral pH and their implications for gene regulation by AraC. A comparison with the structurally unrelated E. coli periplasmic sugar-binding proteins reveals that conserved features of carbohydrate recognition are present, despite a complete lack of structural similarity between the two classes of proteins, suggesting convergent evolution of carbohydrate binding.
- Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205-2185, USA.
Organizational Affiliation: