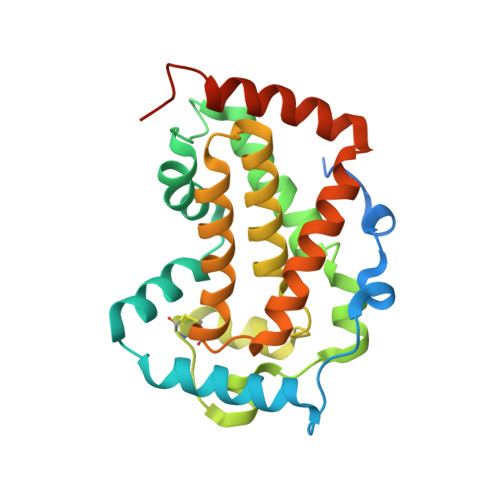
Structure and Mutational Analysis of the PhoN Protein of Salmonella typhimurium Provide Insight into Mechanistic Details.
Makde, R.D., Mahajan, S.K., Kumar, V.(2007) Biochemistry 46: 2079-2090
- PubMed: 17263560 Search on PubMed
- DOI: https://doi.org/10.1021/bi062180g
- Primary Citation Related Structures:
2A96, 2AKC, 2IPB - PubMed Abstract:
The Salmonella typhimurium PhoN protein is a nonspecific acid phosphatase and belongs to the phosphatidic acid phosphatase type 2 (PAP2) superfamily. We report here the crystal structures of phosphate-bound PhoN, the PhoN-tungstate complex, and the T159D mutant of PhoN along with functional characterization of three mutants: L39T, T159D, and D201N. Invariant active site residues, Lys-123, Arg-130, Ser-156, Gly-157, His-158, and Arg-191, interact with phosphate and tungstate oxyanions. Ser-156 also accepts a hydrogen bond from Thr-159. The T159D mutation, surprisingly, severely diminishes phosphatase activity, apparently by disturbing the active site scaffold: Arg-191 is swung out of the active site resulting in conformational changes in His-158 and His-197 residues. Our results reveal a hitherto unknown functional role of Arg-191, namely, restricting the active conformation of catalytic His-158 and His-197 residues. Consistent with the conserved nature of Asp-201 in the PAP2 superfamily, the D201N mutation completely abolished phosphatase activity. On the basis of this observation and in silico analysis we suggest that the crucial mechanistic role of Asp-201 is to stabilize the positive charge on the phosphohistidine intermediate generated by the transfer of phosphoryl to the nucleophile, His-197, located within hydrogen bond distance to the invariant Asp-201. This is in contrast to earlier suggestions that Asp-201 stabilizes His-197 and the His197-Asp201 dyad facilitates formation of the phosphoenzyme intermediate through a charge-relay system. Finally, the L39T mutation in the conserved polyproline motif (39LPPPP43) of dimeric PhoN leads to a marginal reduction in activity, in contrast to the nearly 50-fold reduction observed for monomeric Prevotella intermedia acid phosphatase, suggesting that the varying quaternary structure of PhoN orthologues may have functional significance.
- High Pressure Physics Division, Bhabha Atomic Research Centre, Mumbai 400 085, India.
Organizational Affiliation: