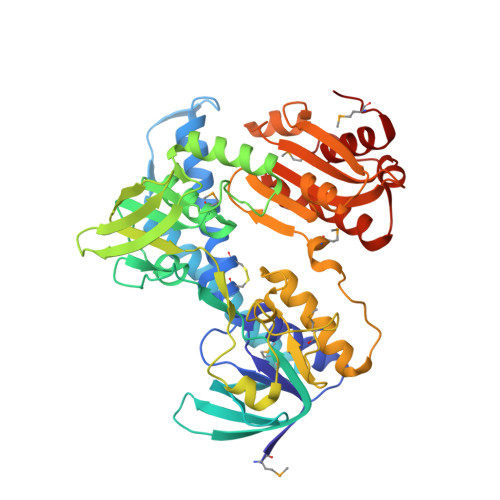
Crystal structure and functional analysis of lipoamide dehydrogenase from Mycobacterium tuberculosis
Rajashankar, K.R., Bryk, R., Kniewel, R., Buglino, J.A., Nathan, C.F., Lima, C.D.(2005) J Biological Chem 280: 33977-33983
- PubMed: 16093239 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M507466200
- Primary Citation Related Structures:
2A8X - PubMed Abstract:
We report the 2.4 A crystal structure for lipoamide dehydrogenase encoded by lpdC from Mycobacterium tuberculosis. Based on the Lpd structure and sequence alignment between bacterial and eukaryotic Lpd sequences, we generated single point mutations in Lpd and assayed the resulting proteins for their ability to catalyze lipoamide reduction/oxidation alone and in complex with other proteins that participate in pyruvate dehydrogenase and peroxidase activities. The results suggest that amino acid residues conserved in mycobacterial species but not conserved in eukaryotic Lpd family members modulate either or both activities and include Arg-93, His-98, Lys-103, and His-386. In addition, Arg-93 and His-386 are involved in forming both "open" and "closed" active site conformations, suggesting that these residues play a role in dynamically regulating Lpd function. Taken together, these data suggest protein surfaces that should be considered while developing strategies for inhibiting this enzyme.
- Structural Biology Program, Sloan-Kettering Institute, New York, New York 10021, USA.
Organizational Affiliation: