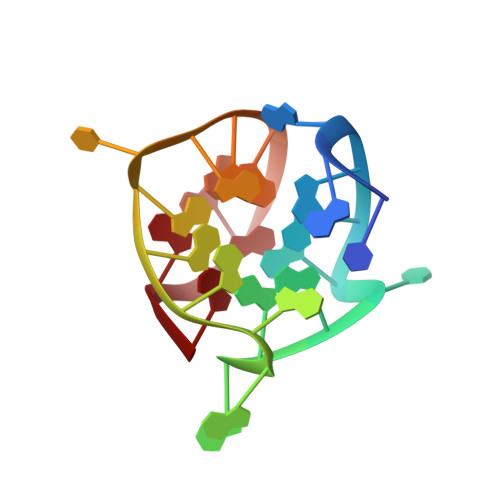
Small-molecule interaction with a five-guanine-tract G-quadruplex structure from the human MYC promoter.
Phan, A.T., Kuryavyi, V., Gaw, H.Y., Patel, D.J.(2005) Nat Chem Biol 1: 167-173
- PubMed: 16408022 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nchembio723
- Primary Citation Related Structures:
2A5P, 2A5R - PubMed Abstract:
It has been widely accepted that DNA can adopt other biologically relevant structures beside the Watson-Crick double helix. One recent important example is the guanine-quadruplex (G-quadruplex) structure formed by guanine tracts found in the MYC (or c-myc) promoter region, which regulates the transcription of the MYC oncogene. Stabilization of this G-quadruplex by ligands, such as the cationic porphyrin TMPyP4, decreases the transcriptional level of MYC. Here, we report the first structure of a DNA fragment containing five guanine tracts from this region. An unusual G-quadruplex fold, which was derived from NMR restraints using unambiguous model-independent resonance assignment approaches, involves a core of three stacked guanine tetrads formed by four parallel guanine tracts with all anti guanines and a snapback 3'-end syn guanine. We have determined the structure of the complex formed between this G-quadruplex and TMPyP4. This structural information, combined with details of small-molecule interaction, provides a platform for the design of anticancer drugs targeting multi-guanine-tract sequences that are found in the MYC and other oncogenic promoters, as well as in telomeres.
- Structural Biology Program, Memorial Sloan-Kettering Cancer Center, New York, New York 10021, USA. phantuan@mskcc.org
Organizational Affiliation: