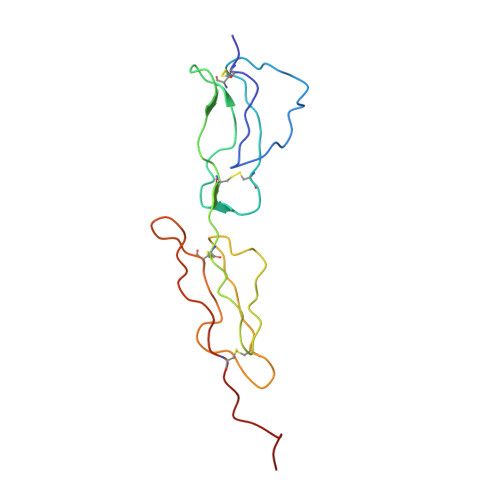
Human C4b-binding Protein, Structural Basis for Interaction with Streptococcal M Protein, a Major Bacterial Virulence Factor
Jenkins, H.T., Mark, L., Ball, G., Persson, J., Lindahl, G., Uhrin, D., Blom, A.M., Barlow, P.N.(2006) J Biological Chem 281: 3690-3697
- PubMed: 16330538 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M511563200
- Primary Citation Related Structures:
2A55 - PubMed Abstract:
Human C4b-binding protein (C4BP) protects host tissue, and those pathogens able to hijack this plasma glycoprotein, from complement-mediated destruction. We now show that the first two complement control protein (CCP) modules of the C4BP alpha-chain, plus the four residues connecting them, are necessary and sufficient for binding a bacterial virulence factor, the Streptococcus pyogenes M4 (Arp4) protein. Structure determination by NMR reveals two tightly coupled CCP modules in an elongated arrangement within this region of C4BP. Chemical shift perturbation studies demonstrate that the N-terminal, hypervariable region of M4 binds to a site including strand 1 of CCP module 2. This interaction is accompanied by an intermodular reorientation within C4BP. We thus provide a detailed picture of an interaction whereby a pathogen evades complement.
- Edinburgh Biological NMR Unit, University of Edinburgh, West Mains Road, Edinburgh EH9 3JJ, Scotland, United Kingdom.
Organizational Affiliation: