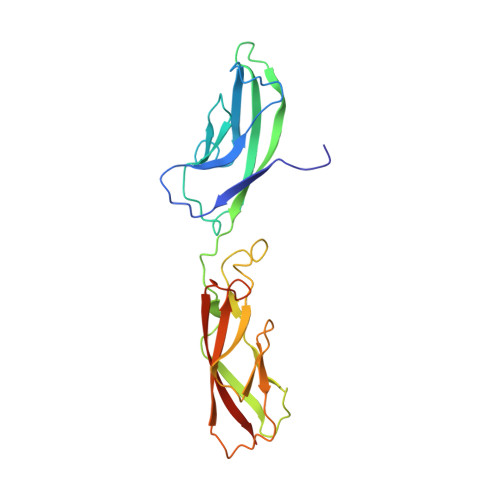
Type II cadherin ectodomain structures: implications for classical cadherin specificity.
Patel, S.D., Ciatto, C., Chen, C.P., Bahna, F., Rajebhosale, M., Arkus, N., Schieren, I., Jessell, T.M., Honig, B., Price, S.R., Shapiro, L.(2006) Cell 124: 1255-1268
- PubMed: 16564015 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2005.12.046
- Primary Citation Related Structures:
1ZVN, 1ZXK, 2A4C, 2A4E, 2A62 - PubMed Abstract:
Type I and II classical cadherins help to determine the adhesive specificities of animal cells. Crystal-structure determination of ectodomain regions from three type II cadherins reveals adhesive dimers formed by exchange of N-terminal beta strands between partner extracellular cadherin-1 (EC1) domains. These interfaces have two conserved tryptophan side chains that anchor each swapped strand, compared with one in type I cadherins, and include large hydrophobic regions unique to type II interfaces. The EC1 domains of type I and type II cadherins appear to encode cell adhesive specificity in vitro. Moreover, perturbation of motor neuron segregation with chimeric cadherins depends on EC1 domain identity, suggesting that this region, which includes the structurally defined adhesive interface, encodes type II cadherin functional specificity in vivo.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: