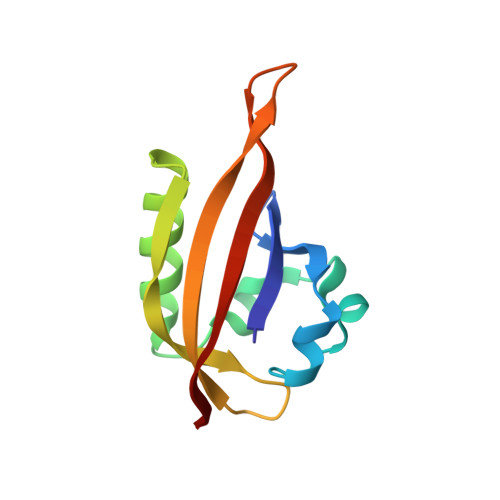
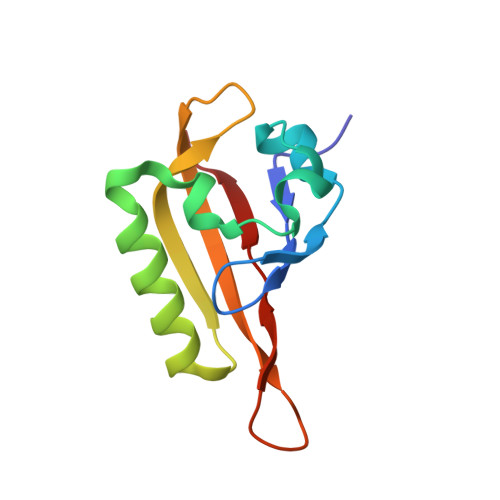
Structural basis of ARNT PAS-B dimerization: use of a common beta-sheet interface for hetero- and homodimerization.
Card, P.B., Erbel, P.J., Gardner, K.H.(2005) J Mol Biology 353: 664-677
- PubMed: 16181639 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.08.043
- Primary Citation Related Structures:
1X0O, 2A24 - PubMed Abstract:
The aryl hydrocarbon receptor nuclear translocator (ARNT) is a promiscuous bHLH-PAS (Per-ARNT-Sim) protein that forms heterodimeric transcriptional regulator complexes with several other bHLH-PAS subunits to control a variety of biological pathways, some of which are centrally involved in disease initiation and/or progression. One of these is the hypoxia response pathway, which allows eukaryotic cells to respond to low oxygen tension via the formation of a heterodimeric complex between ARNT and another bHLH-PAS protein, the hypoxia-inducible factor alpha (HIF-alpha). We have previously shown that the C-terminal PAS domains of an HIF-alpha isoform (HIF-2alpha) and ARNT interact in vitro, and that mutations in the solvent-exposed beta-sheet surface of the HIF-2alpha domain not only disrupt this interaction, but also greatly attenuate the hypoxia response in living cells. Here, we have solved the solution structure of the corresponding PAS domain of ARNT and show that it utilizes a very similar interface for the interaction with the HIF-2alpha PAS domain. We also show that this domain self-associates in a concentration-dependent manner, and that the interface used in this homodimeric complex is very similar to that used in the formation of heterodimer. In addition, using experimentally derived NMR restraints, we used the program HADDOCK to calculate a low-resolution model of the complex formed in solution by these two PAS domains, and confirm the validity of this model using site-directed spin labeling to obtain long-range distance information in solution. With this information, we propose a model for the mode of multi-PAS domain interaction in bHLH-PAS transcriptional activation complexes.
- Department of Biochemistry, UT Southwestern Medical Center, 5323 Harry Hines Blvd., Dallas, TX 75390-8816, USA.
Organizational Affiliation: