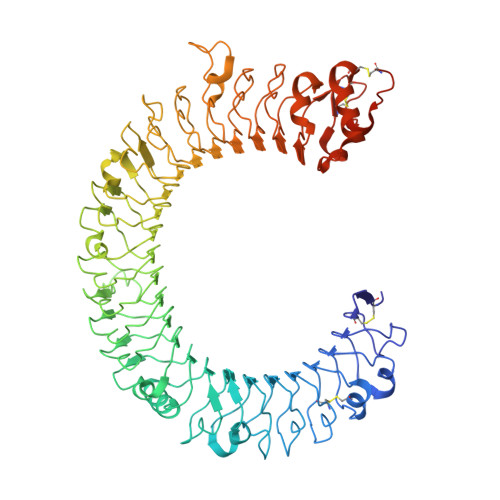
The molecular structure of the Toll-like receptor 3 ligand-binding domain
Bell, J.K., Botos, I., Hall, P.R., Askins, J., Shiloach, J., Segal, D.M., Davies, D.R.(2005) Proc Natl Acad Sci U S A 102: 10976-10980
- PubMed: 16043704 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0505077102
- Primary Citation Related Structures:
2A0Z - PubMed Abstract:
Innate immunity is the first line of defense against invading pathogens. Toll-like receptors (TLRs) act as sentinels of the innate immune system, sensing a variety of ligands from lipopolysaccharide to flagellin to dsRNA through their ligand-binding domain that is composed of leucine-rich repeats (LRRs). Ligand binding initiates a signaling cascade that leads to the up-regulation of inflammation mediators. In this study, we have expressed and crystallized the ectodomain (ECD) of human TLR3, which recognizes dsRNA, a molecular signature of viruses, and have determined the molecular structure to 2.4-A resolution. The overall horseshoe-shaped structure of the TLR3-ECD is formed by 23 repeating LRRs that are capped at each end by specialized non-LRR domains. The extensive beta-sheet on the molecule's concave surface forms a platform for several modifications, including insertions in the LRRs and 11 N-linked glycans. The TLR3-ECD structure indicates how LRR loops can establish distinct pathogen recognition receptors.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: