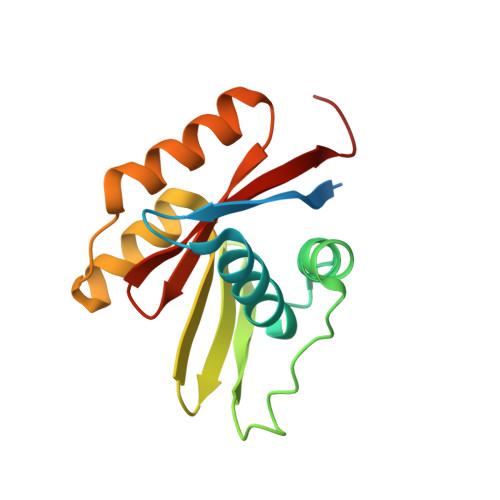
The Holo-form of the Nucleotide Binding Domain of the KdpFABC Complex from Escherichia coli Reveals a New Binding Mode
Haupt, M., Bramkamp, M., Heller, M., Coles, M., Deckers-Hebestreit, G., Herkenhoff-Hesselmann, B., Altendorf, K., Kessler, H.(2006) J Biological Chem 281: 9641-9649
- PubMed: 16354672
- DOI: https://doi.org/10.1074/jbc.M508290200
- Primary Citation of Related Structures:
2A00, 2A29 - PubMed Abstract:
P-type ATPases are ubiquitously abundant enzymes involved in active transport of charged residues across biological membranes. The KdpB subunit of the prokaryotic Kdp-ATPase (KdpFABC complex) shares characteristic regions of homology with class II-IV P-type ATPases and has been shown previously to be misgrouped as a class IA P-type ATPase. Here, we present the NMR structure of the AMP-PNP-bound nucleotide binding domain KdpBN of the Escherichia coli Kdp-ATPase at high resolution. The aromatic moiety of the nucleotide is clipped into the binding pocket by Phe(377) and Lys(395) via a pi-pi stacking and a cation-pi interaction, respectively. Charged residues at the outer rim of the binding pocket (Arg(317), Arg(382), Asp(399), and Glu(348)) stabilize and direct the triphosphate group via electrostatic attraction and repulsion toward the phosphorylation domain. The nucleotide binding mode was corroborated by the replacement of critical residues. The conservative mutation F377Y produced a high residual nucleotide binding capacity, whereas replacement by alanine resulted in low nucleotide binding capacities and a considerable loss of ATPase activity. Similarly, mutation K395A resulted in loss of ATPase activity and nucleotide binding affinity, even though the protein was properly folded. We present a schematic model of the nucleotide binding mode that allows for both high selectivity and a low nucleotide binding constant, necessary for the fast and effective turnover rate realized in the reaction cycle of the Kdp-ATPase.
- Department Chemie, Technische Universität München, Lichtenbergstrasse 4, 85747 Garching, Germany.
Organizational Affiliation: