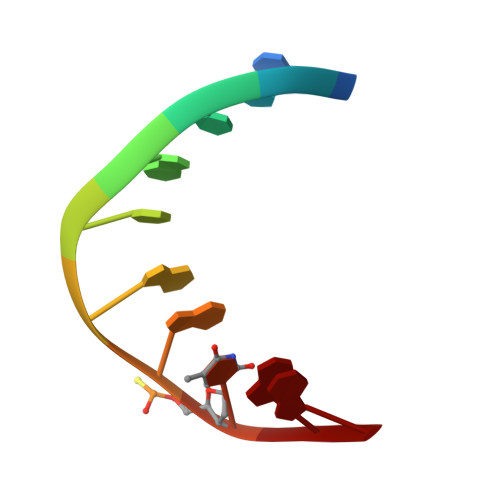
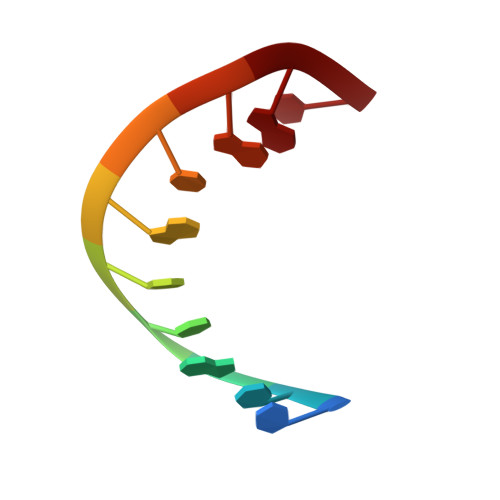
Structure and dynamics of a DNA.RNA hybrid duplex with a chiral phosphorothioate moiety: NMR and molecular dynamics with conventional and time-averaged restraints.
Gonzalez, C., Stec, W., Reynolds, M.A., James, T.L.(1995) Biochemistry 34: 4969-4982
- PubMed: 7711019 Search on PubMed
- DOI: https://doi.org/10.1021/bi00015a008
- Primary Citation Related Structures:
219D - PubMed Abstract:
The three-dimensional structure of two thiophosphate-modified DNA.RNA hybrid duplexes d(GCTATAApsTGG).r(CCAUUAUAGC), one with R-thiophosphate chirality and one with S-thiophosphate chirality, have been determined by restrained molecular dynamics simulations (rMD). As the two yielded almost identical results, a description of results can be presented in the singular. The conformational flexibility of this hybrid has been investigated by employing time-averaged constraints during the molecular dynamics simulations (MD-tar). A set of structural restraints, comprising 322 precise interproton distance constraints obtained by a complete relaxation matrix analysis of the 2D NOE intensities as well as J coupling constants obtained from quantitative simulations of DQF-COSY cross-peaks in deoxyriboses, was reported in our previous paper [González, C., Stec, W., Kobylanska, A., Hogrefe, R. I., Reynolds, M., & James, T. L. (1994) Biochemistry 33, 11062-11072]. Multiple conformations of the deoxyribose moieties were evident from the scalar coupling constant analysis. Accurate distance constraints, obtained from complete relaxation matrix analysis, yielded a time-averaged solution structure via conventional restrained molecular dynamics which is not compatible with the experimental J coupling constants (root-mean-square deviation in J value approximately 2 Hz). However, vicinal coupling constant information can be reproduced when time-averaged constraints are used during the molecular dynamics calculations instead of the conventional restraints (Jrms approximately 0.6 Hz). MD-tar simulations also improve the NMR R factors. This improvement is more evident in the DNA than in the RNA strand, where no indication of conformational flexibility had been obtained. Analysis of the MD-tar trajectories confirms that deoxyriboses undergo pucker transitions between the S and N domain, with the major conformer in the S domain. The ribose moieties in the RNA strand, however, remain in the N domain during the entire simulation. Conformations of deoxyriboses in the intermediate domain near O4'-endo are obtained when the average structure is calculated with conventional NMR restraints. Since these conformations cannot account for the experimental J coupling information, and they only appear in a very low population in the MD-tar ensemble, we conclude that intermediate E sugar puckers are artifacts produced by the attempt to fit all the structural constraints simultaneously when in reality more than one conformer is present. Most structural features of the duplex remain the same in the average structure and in the MD-tar ensemble, e.g., the minor groove width, exhibiting an intermediate value compared with those of canonical A- and B-like structures.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Pharmaceutical Chemistry, University of California, San Francisco 94143-0446, USA.
Organizational Affiliation: