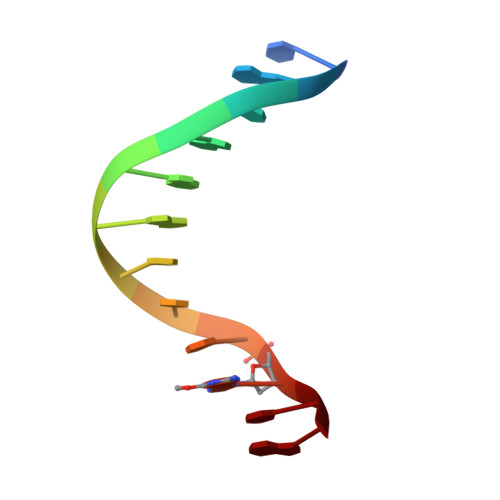
Structure of a new crystal form of a DNA dodecamer containing T.(O6Me)G base pairs.
Vojtechovsky, J., Eaton, M.D., Gaffney, B., Jones, R., Berman, H.M.(1995) Biochemistry 34: 16632-16640
- PubMed: 8527436 Search on PubMed
- DOI: https://doi.org/10.1021/bi00051a011
- Primary Citation Related Structures:
218D - PubMed Abstract:
The structure of the synthetic dodecamer (d[CGTGAATTC(O6Me)GCG])2 has been determined to a resolution of 2.25 A and refined to a final R factor of 16.7%. The volume of the unit cell is significantly smaller by 16% than the original Drew and Dickerson parent dodecamer [Drew, H. R., Wing, R. M., Takano, T., Broka, C., Tanaka, S., Itakura, K., & Dickerson, R. E. (1981) Proc. Natl. Acad. Sci. U.S.A. 78, 7318-7322]. The double helix is in a different position in the unit cell, rotated by -85.9 degrees, and translated by 9.9 A around the helical axis with respect to the parent structure. The intermolecular arrangement of helices, characterized by double hydrogen bonded guanine-guanine minor groove interactions, remains conserved. The molecular geometry exhibits several significant changes that are related to the changed position of the helix and the presence of two mismatched base pairs with O6-methylguanine. Both mispairs are found in a symmetrical T(anti).(O6Me)G(anti) conformation, and the methyl groups are in proximal orientation. The hydration pattern of the structure is different and can be related to changes in the minor groove geometry. An incorrect model that was isomorphous to the parent dodecamer could be refined to a low R factor. Characteristics of the refinement and of the geometry that are indicative of incorrect structures have been analyzed.
- Department of Chemistry, Rutgers University, Piscataway, New Jersey 08855-0939, USA.
Organizational Affiliation: