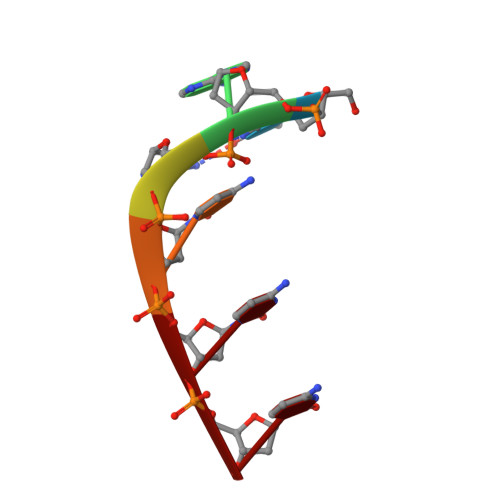
Stable loop in the crystal structure of the intercalated four-stranded cytosine-rich metazoan telomere.
Kang, C., Berger, I., Lockshin, C., Ratliff, R., Moyzis, R., Rich, A.(1995) Proc Natl Acad Sci U S A 92: 3874-3878
- PubMed: 7731999 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.92.9.3874
- Primary Citation Related Structures:
200D - PubMed Abstract:
In most metazoans, the telomeric cytosine-rich strand repeating sequence is d(TAACCC). The crystal structure of this sequence was solved to 1.9-A resolution. Four strands associate via the cytosine-containing parts to form a four-stranded intercalated structure held together by C.C+ hydrogen bonds. The base-paired strands are parallel to each other, and the two duplexes are intercalated into each other in opposite orientations. One TAA end forms a highly stabilized loop with the 5' thymine Hoogsteen-base-paired to the third adenine. The 5' end of this loop is in close proximity to the 3' end of one of the other intercalated cytosine strands. Instead of being entirely in a DNA duplex, this structure suggests the possibility of an alternative conformation for the cytosine-rich telomere strands.
- Department of Biology, Massachusetts Institute of Technology, Cambridge 02139, USA.
Organizational Affiliation: