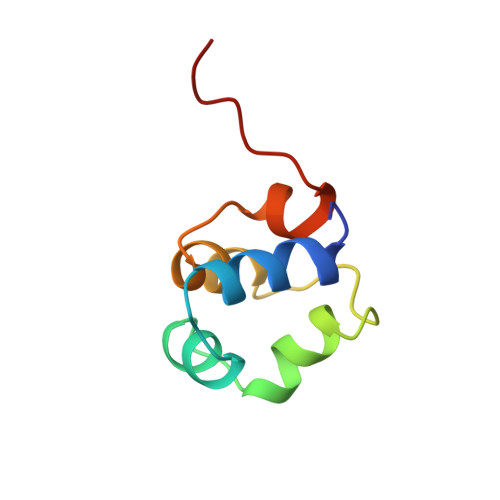
Three-dimensional solution structure and stability of phage 434 Cro protein.
Padmanabhan, S., Jimenez, M.A., Gonzalez, C., Sanz, J.M., Gimenez-Gallego, G., Rico, M.(1997) Biochemistry 36: 6424-6436
- PubMed: 9174359
- DOI: https://doi.org/10.1021/bi970085p
- Primary Citation of Related Structures:
1ZUG - PubMed Abstract:
1H NMR resonances of the phage 434 Cro protein were assigned using standard 2D NMR methods, and its solution structure determined using 867 distance constraints in distance geometry (DIANA) calculations ultimately refined by restrained molecular dynamics (GROMOS). In the 20 best NMR structures, the average pairwise backbone and heavy atom RMSDs are 0.63 +/- 0.14 and 1.53 +/- 0.15 A, respectively, for the structurally well-defined residues 4-65. Residues 1-3 and 66-71 at the N- and C-termini are structurally disordered. The region 4-65 includes five alpha-helices and tight turns which define the hydrophobic core of the protein. The backbone and heavy atom RMSDs for residues 4-65 are 0.92 +/- 0.12 and 1.99 +/- 0.12 A, respectively, for the NMR versus the crystal structures, but there are significant differences in the side-chain conformations and solvent accessibilities for some core residues. Analytical ultracentrifugation experiments confirm that 434 Cro is monomeric even at the high NMR concentrations. 434 Cro folding under NMR solution conditions is two-state as indicated by coincident urea denaturation curves from circular dichroism and intrinsic fluorescence measurements. They yield values for 434 Cro stability which show good correspondence to the free energy for global unfolding determined by NMR hydrogen exchange measurements for the slowest exchanging amide protons.
- Instituto de Estructura de la Materia, Consejo Superior de Investigaciones Cientificas, Madrid, Spain.
Organizational Affiliation: