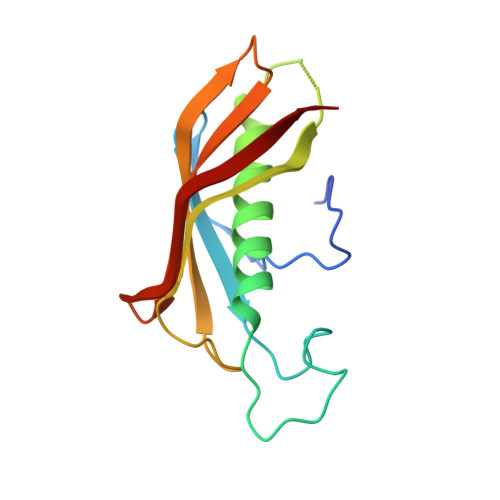
Crystal structure of dimeric FabZ of Plasmodium falciparum reveals conformational switching to active hexamers by peptide flips
Swarnamukhi, P.L., Sharma, S.K., Bajaj, P., Surolia, N., Surolia, A., Suguna, K.(2006) FEBS Lett 580: 2653-2660
- PubMed: 16643907
- DOI: https://doi.org/10.1016/j.febslet.2006.04.014
- Primary Citation of Related Structures:
1ZHG - PubMed Abstract:
The crystal structure of beta-hydroxyacyl acyl carrier protein dehydratase of Plasmodium falciparum (PfFabZ) has been determined at a resolution of 2.4 A. PfFabZ has been found to exist as a homodimer (d-PfFabZ) in the crystals of the present study in contrast to the reported hexameric form (h-PfFabZ) which is a trimer of dimers crystallized in a different condition. The catalytic sites of this enzyme are located in deep narrow tunnel-shaped pockets formed at the dimer interface. A histidine residue from one subunit of the dimer and a glutamate residue from the other subunit lining the tunnel form the catalytic dyad in the reported crystal structures. While the position of glutamate remains unaltered in the crystal structure of d-PfFabZ compared to that in h-PfFabZ, the histidine residue takes up an entirely different conformation and moves away from the tunnel leading to a His-Phe cis-trans peptide flip at the histidine residue. In addition, a loop in the vicinity has been observed to undergo a similar flip at a Tyr-Pro peptide bond. These alterations not only prevent the formation of a hexamer but also distort the active site geometry resulting in a dimeric form of FabZ that is incapable of substrate binding. The dimeric state and an altered catalytic site architecture make d-PfFabZ distinctly different from the FabZ structures described so far. Dynamic light scattering and size exclusion chromatographic studies clearly indicate a pH-related switching of the dimers to active hexamers.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore 560 012, India.
Organizational Affiliation: