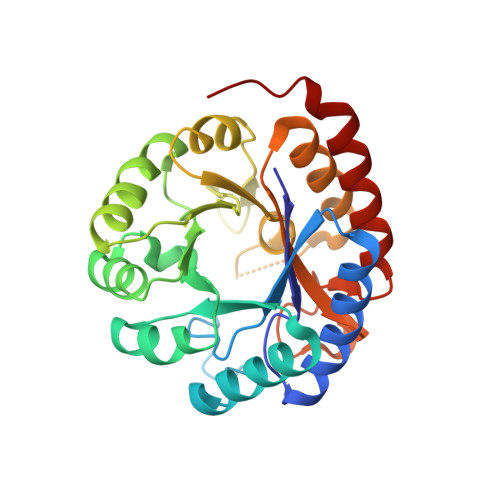
The Catalytic and Conformational Cycle of Aquifex aeolicus KDO8P Synthase: Role of the L7 Loop.
Xu, X., Kona, F., Wang, J., Lu, J., Stemmler, T., Gatti, D.L.(2005) Biochemistry 44: 12434-12444
- PubMed: 16156656
- DOI: https://doi.org/10.1021/bi051095q
- Primary Citation Related Structures:
1ZHA, 1ZJI - PubMed Abstract:
KDO8P synthase catalyzes the condensation of arabinose 5-phosphate (A5P) and phosphoenolpyruvate (PEP) to form the 8-carbon sugar KDO8P and inorganic phosphate (P(i)). The X-ray structure of the wild-type enzyme shows that when both PEP and A5P bind, the active site becomes isolated from the environment due to a conformational change of the L7 loop. The structures of the R106G mutant, without substrates, and with PEP and PEP plus A5P bound, were determined and reveal that in R106G closure of the L7 loop is impaired. The structural perturbations originating from the loss of the Arg(106) side chain point to a role of the L2 loop in stabilizing the closed conformation of the L7 loop. Despite the increased exposure of the R106G active site, no abnormal reaction of PEP with water was observed, ruling out the hypothesis that the primary function of the L7 loop is to shield the active site from bulk solvent during the condensation reaction. However, the R106G enzyme displays several kinetic abnormalities on both the substrate side (smaller K(m)(PEP), larger K(i)(A5P) and K(m)(A5P)) and the product side (smaller K(i)(Pi) and K(i)(KDO8P)) of the reaction. As a consequence, the mutant enzyme is less severely inhibited by A5P and more severely inhibited by P(i) and KDO8P. Simulations of the flux of KDO8P synthesis under metabolic steady-state conditions (constant concentration of reactants and products over time) suggest that in vivo R106G is expected to perform optimally in a narrower range of substrate and product concentrations than the wild-type enzyme.
- Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, Michigan 48201, USA.
Organizational Affiliation: