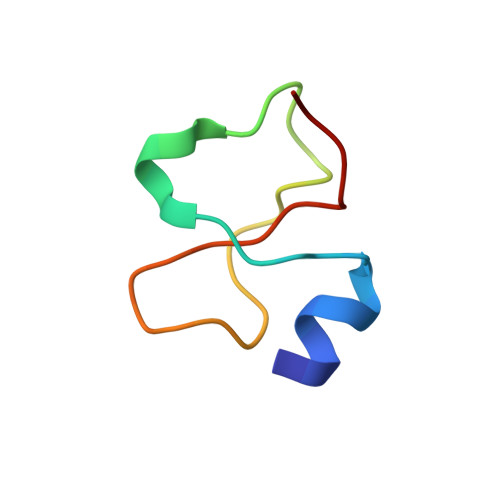
Automated NMR structure determination and disulfide bond identification of the myotoxin crotamine from Crotalus durissus terrificus.
Fadel, V., Bettendorff, P., Herrmann, T., de Azevedo, W.F., Oliveira, E.B., Yamane, T., Wuthrich, K.(2005) Toxicon 46: 759-767
- PubMed: 16185738 Search on PubMed
- DOI: https://doi.org/10.1016/j.toxicon.2005.07.018
- Primary Citation Related Structures:
1Z99 - PubMed Abstract:
Crotamine is one of four major components of the venom of the South American rattlesnake Crotalus durissus terrificus. Similar to its counterparts in the family of the myotoxins, it induces myonecrosis of skeletal muscle cells. This paper describes a new NMR structure determination of crotamine in aqueous solution at pH 5.8 and 20 degrees C, using standard homonuclear 1H NMR spectroscopy at 900MHz and the automated structure calculation software ATNOS/CANDID/DYANA. The automatic NOESY spectral analysis included the identification of a most likely combination of the six cysteines into three disulfide bonds, i.e. Cys4-Cys36, Cys11-Cys30 and Cys18-Cys37; thereby a generally applicable new computational protocol is introduced to determine unknown disulfide bond connectivities in globular proteins. A previous NMR structure determination was thus confirmed and the structure refined. Crotamine contains an alpha-helix with residues 1-7 and a two-stranded anti-parallel beta-sheet with residues 9-13 and 34-38 as the only regular secondary structures. These are connected with each other and the remainder of the polypeptide chain by the three disulfide bonds, which also form part of a central hydrophobic core. A single conformation was observed, with Pro13 and Pro21 in the trans and Pro20 in the cis-form. The global fold and the cysteine-pairing pattern of crotamine are similar to the beta-defensin fold, although the two proteins have low sequence homology, and display different biological activities.
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule Zürich, CH-8093 Zurich, Switzerland.
Organizational Affiliation: