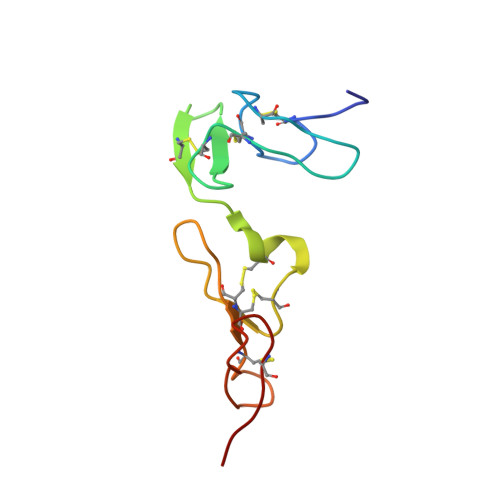
Solution Structure of the Ca(2+)-Binding EGF3-4 Pair from Vitamin K-Dependent Protein S: Identification of an Unusual Fold in EGF3.
Drakenberg, T., Ghasriani, H., Thulin, E., Muranyi, A., Annila, A., Stenflo, J.(2005) Biochemistry 44: 8782-8789
- PubMed: 15952784 Search on PubMed
- DOI: https://doi.org/10.1021/bi050101f
- Primary Citation Related Structures:
1Z6C - PubMed Abstract:
Vitamin K-dependent protein S is a cofactor of activated protein C, a serine protease that regulates blood coagulation. Deficiency of protein S can cause venous thrombosis. Protein S has four EGF domains in tandem; domains 2-4 bind calcium with high affinity whereas domains 1-2 mediate interaction with activated protein C. We have now solved the solution structure of the EGF3-4 fragment of protein S. The linker between the two domains is similar to what has been observed in other calcium-binding EGF domains where it provides an extended conformation. Interestingly, a disagreement between NOE and RDC data revealed a conformational heterogeneity within EGF3 due to a hinge-like motion around Glu186 in the Cys-Glu-Cys sequence, the only point in the domain where flexibility is allowed. The dominant, bent conformation of EGF3 in the pair has no precedent among calcium-binding EGF domains. It is characterized by a change in the psi angle of Glu186 from 160 degrees +/- 40 degrees , as seen in ten other EGF domains, to approximately 0 degrees +/- 15 degrees . NOESY data suggest that Tyr193, a residue not conserved in other calcium-binding EGF domains (except in the homologue Gas6), induces the unique fold of EGF3. However, SAXS data, obtained on EGF1-4 and EGF2-4, showed a dominant, extended conformation in these fragments. This may be due to a counterproductive domain-domain interaction between EGF2 and EGF4 if EGF3 is in a bent conformation. We speculate that the ability of EGF3 to adopt different conformations may be of functional significance in protein-protein interactions involving protein S.
- Department of Biophysical Chemistry, University of Lund, P.O. Box 124, SE-221 00 Lund, Sweden. torbjorn.drakenberg@bpc.lu.se
Organizational Affiliation: