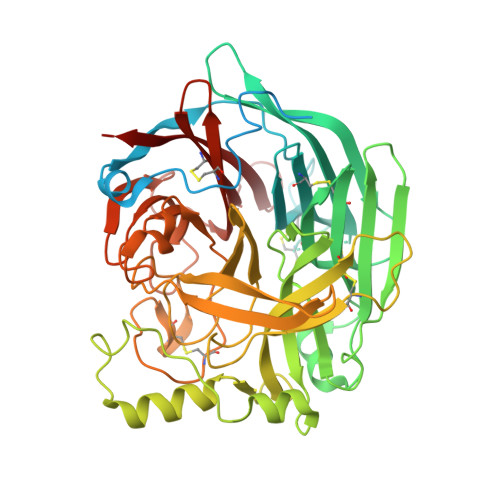
Structural studies of the parainfluenza virus 5 hemagglutinin-neuraminidase tetramer in complex with its receptor, sialyllactose.
Yuan, P., Thompson, T.B., Wurzburg, B.A., Paterson, R.G., Lamb, R.A., Jardetzky, T.S.(2005) Structure 13: 803-815
- PubMed: 15893670 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.02.019
- Primary Citation Related Structures:
1Z4V, 1Z4W, 1Z4X, 1Z4Y, 1Z4Z, 1Z50 - PubMed Abstract:
The paramyxovirus hemagglutinin-neuraminidase (HN) functions in virus attachment to cells, cleavage of sialic acid from oligosaccharides, and stimulating membrane fusion during virus entry into cells. The structural basis for these diverse functions remains to be fully understood. We report the crystal structures of the parainfluenza virus 5 (SV5) HN and its complexes with sialic acid, the inhibitor DANA, and the receptor sialyllactose. SV5 HN shares common structural features with HN of Newcastle disease virus (NDV) and human parainfluenza 3 (HPIV3), but unlike the previously determined HN structures, the SV5 HN forms a tetramer in solution, which is thought to be the physiological oligomer. The sialyllactose complex reveals intact receptor within the active site, but no major conformational changes in the protein. The SV5 HN structures do not support previously proposed models for HN action in membrane fusion and suggest alternative mechanisms by which HN may promote virus entry into cells.
- Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois 60208-3500, USA.
Organizational Affiliation: