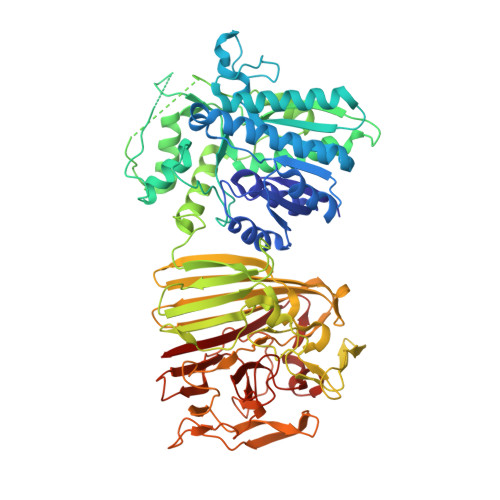
The molecular architecture of galactose mutarotase/UDP-galactose 4-epimerase from Saccharomyces cerevisiae.
Thoden, J.B., Holden, H.M.(2005) J Biological Chem 280: 21900-21907
- PubMed: 15795221 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M502411200
- Primary Citation Related Structures:
1Z45 - PubMed Abstract:
The metabolic pathway by which beta-D-galactose is converted to glucose 1-phosphate is known as the Leloir pathway and consists of four enzymes. In most organisms, these enzymes appear to exist as soluble entities in the cytoplasm. In yeast such as Saccharomyces cerevisiae, however, the first and last enzymes of the pathway, galactose mutarotase and UDP-galactose 4-epimerase, are contained within a single polypeptide chain referred to as Gal10p. Here we report the three-dimensional structure of Gal10p in complex with NAD(+), UDP-glucose, and beta-D-galactose determined to 1.85-A resolution. The enzyme is dimeric with dimensions of approximately 91 A x 135 A x 108 A and assumes an almost V-shaped appearance. The overall architecture of the individual subunits can be described in terms of two separate N- and C-terminal domains connected by a Type II turn formed by Leu-357 to Val-360. The first 356 residues of Gal10p fold into the classical bilobal topology observed for all other UDP-galactose 4-epimerases studied thus far. This N-terminal domain contains the binding sites for NAD(+) and UDP-glucose. The polypeptide chain extending from Glu-361 to Ser-699 adopts a beta-sandwich motif and harbors the binding site for beta-D-galactose. The two active sites of Gal10p are separated by over 50 A. This investigation represents the first structural analysis of a dual function enzyme in the Leloir pathway.
- Department of Biochemistry, University of Wisconsin, Madison, WI 53706, USA.
Organizational Affiliation: