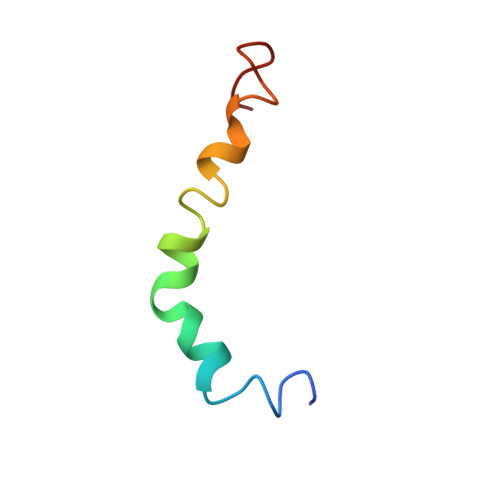
The alpha-to-beta Conformational Transition of Alzheimer's Abeta-(1-42) Peptide in Aqueous Media is Reversible: A Step by Step Conformational Analysis Suggests the Location of beta Conformation Seeding
Tomaselli, S., Esposito, V., Vangone, P., van Nuland, N.A., Bonvin, A.M., Guerrini, R., Tancredi, T., Temussi, P.A., Picone, D.(2006) Chembiochem 7: 257-267
- PubMed: 16444756 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.200500223
- Primary Citation Related Structures:
1Z0Q - PubMed Abstract:
Current views of the role of beta-amyloid (Abeta) peptide fibrils range from regarding them as the cause of Alzheimer's pathology to having a protective function. In the last few years, it has also been suggested that soluble oligomers might be the most important toxic species. In all cases, the study of the conformational properties of Abeta peptides in soluble form constitutes a basic approach to the design of molecules with "antiamyloid" activity. We have experimentally investigated the conformational path that can lead the Abeta-(1-42) peptide from the native state, which is represented by an alpha helix embedded in the membrane, to the final state in the amyloid fibrils, which is characterized by beta-sheet structures. The conformational steps were monitored by using CD and NMR spectroscopy in media of varying polarities. This was achieved by changing the composition of water and hexafluoroisopropanol (HFIP). In the presence of HFIP, beta conformations can be observed in solutions that have very high water content (up to 99 % water; v/v). These can be turned back to alpha helices simply by adding the appropriate amount of HFIP. The transition of Abeta-(1-42) from alpha to beta conformations occurs when the amount of water is higher than 80 % (v/v). The NMR structure solved in HFIP/H2O with high water content showed that, on going from very apolar to polar environments, the long N-terminal helix is essentially retained, whereas the shorter C-terminal helix is lost. The complete conformational path was investigated in detail with the aid of molecular-dynamics simulations in explicit solvent, which led to the localization of residues that might seed beta conformations. The structures obtained might help to find regions that are more affected by environmental conditions in vivo. This could in turn aid the design of molecules able to inhibit fibril deposition or revert oligomerization processes.
- Dipartimento di Chimica, Università Federico II di Napoli, Via Cintia, 80126 Napoli, Italy.
Organizational Affiliation: