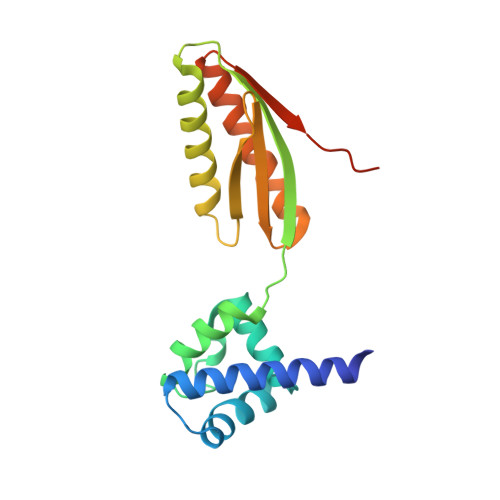
Structure-Function Relationships of the Intact aIF2alpha Subunit from the Archaeon Pyrococcus abyssi
Yatime, L., Schmitt, E., Blanquet, S., Mechulam, Y.(2005) Biochemistry 44: 8749-8756
- PubMed: 15952781
- DOI: https://doi.org/10.1021/bi050373i
- Primary Citation of Related Structures:
1YZ6, 1YZ7 - PubMed Abstract:
Eukaryotic and archaeal initiation factor 2 (e- and aIF2, respectively) are heterotrimeric proteins (alphabetagamma) supplying the small subunit of the ribosome with methionylated initiator tRNA. The gamma subunit forms the core of the heterotrimer. It resembles elongation factor EF1-A and ensures interaction with Met-tRNA(i)(Met). In the presence of the alpha subunit, which is composed of three domains, the gamma subunit expresses full tRNA binding capacity. This study reports the crystallographic structure of the intact aIF2alpha subunit from the archaeon Pyrococcus abyssi and that of a derived C-terminal fragment containing domains 2 and 3. The obtained structures are compared with those of N-terminal domains 1 and 2 of yeast and human eIF2alpha and with the recently determined NMR structure of human eIF2alpha. We show that the three-domain organization in the alpha subunit is conserved in archaea and eukarya. Domains 1 and 2 form a rigid body linked to a mobile third domain. Sequence comparisons establish that the most conserved regions in the aIF2alpha polypeptide lie at opposite sides of the protein, within domain 1 and domain 3, respectively. These two domains are known to exhibit RNA binding capacities. We propose that domain 3, which is known to glue the alpha subunit onto the gamma subunit, participates in Met-tRNA(i)(Met) binding while domain 1 recognizes either rRNA or mRNA on the ribosome. Thereby, the observed structural mobility within the e- and aIF2alpha molecules would be an integral part of the biological function of this subunit in the heterotrimeric e- and aIF2alphabetagamma factors.
- Laboratoire de Biochimie, Unité Mixte de Recherche 7654, CNRS-Ecole Polytechnique, F-91128 Palaiseau Cedex, France.
Organizational Affiliation: