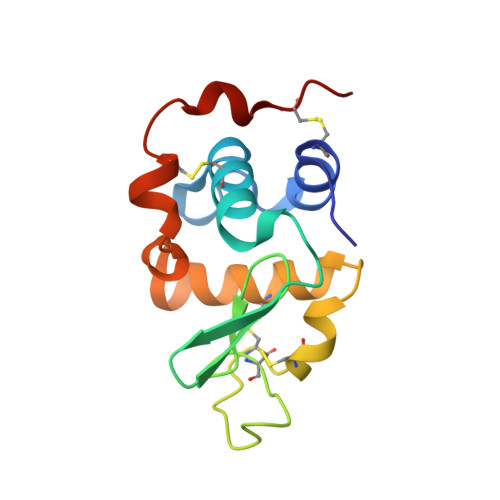
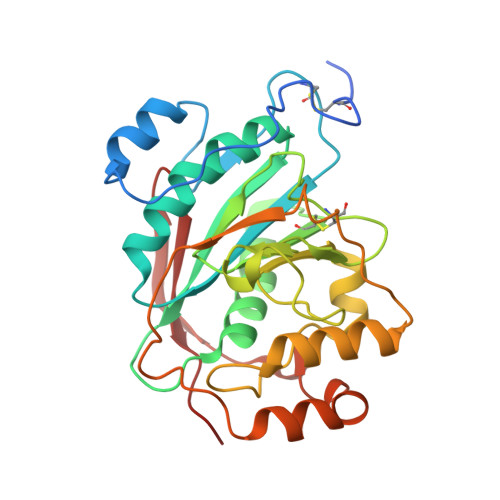
Mutation of Arginine 228 to Lysine Enhances the Glucosyltransferase Activity of Bovine beta-1,4-Galactosyltransferase I
Ramakrishnan, B., Boeggeman, E., Qasba, P.K.(2005) Biochemistry 44: 3202-3210
- PubMed: 15736931 Search on PubMed
- DOI: https://doi.org/10.1021/bi0479454
- Primary Citation Related Structures:
1YRO - PubMed Abstract:
Beta-1,4-galactosyltransferase I (beta4Gal-T1) normally transfers Gal from UDP-Gal to GlcNAc in the presence of Mn(2+) ion (Gal-T activity) and also transfers Glc from UDP-Glc to GlcNAc (Glc-T activity), albeit at only 0.3% efficiency. In addition, alpha-lactalbumin (LA) enhances this Glc-T activity more than 25 times. Comparison of the crystal structures of UDP-Gal- and UDP-Glc-bound beta4Gal-T1 reveals that the O4 hydroxyl group in both Gal and Glc moieties forms a hydrogen bond with the side chain carboxylate group of Glu317. The orientation of the O4 hydroxyl of glucose causes a steric hindrance to the side chain carboxylate group of Glu317, accounting for the enzyme's low Glc-T activity. In this study, we show that mutation of Arg228, a residue in the vicinity of Glu317, to lysine (R228K-Gal-T1) results in a 15-fold higher Glc-T activity, which is further enhanced by LA to nearly 25% of the Gal-T activity of the wild type. The kinetic parameters indicate that the main effect of the mutation of Arg228 to lysine is on the k(cat) of Glc-T, which increases 3-4-fold, both in the absence and in the presence of LA; simultaneously, the k(cat) for the Gal-T reaction is reduced 30-fold. The crystal structure of R228K-Gal-T1 complexed with LA, UDP-Gal, and Mn(2+) determined at 1.9 A resolution shows that the Asp318 side chain exhibits a minor alternate conformation, compared to that in the wild type. This alternate conformation now causes a steric hindrance to the O4 hydroxyl group of the Gal moiety of UDP-Gal, probably causing the dissociation of UDP-Gal and the reduced k(cat) of the Gal-T reaction.
- Structural Glycobiology Section, Laboratory of Experimental and Computational Biology, Center for Cancer Research, National Cancer Institute-Frederick, Frederick, Maryland 21702-1201, USA.
Organizational Affiliation: