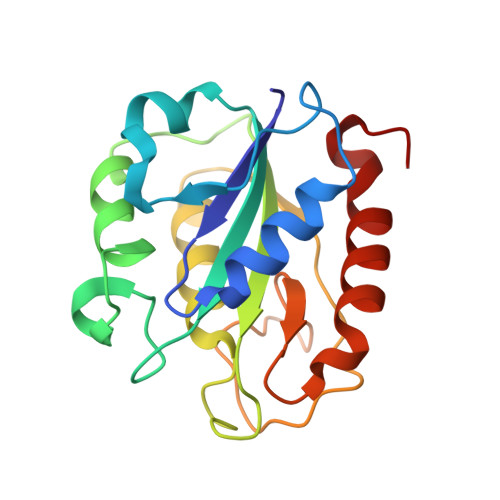
A crystallographic study of Cys69Ala flavodoxin II from Azotobacter vinelandii: structural determinants of redox potential
Alagaratnam, S., van Pouderoyen, G., Pijning, T., Dijkstra, B.W., Cavazzini, D., Rossi, G.L., Van Dongen, W.M., van Mierlo, C.P., van Berkel, W.J., Canters, G.W.(2005) Protein Sci 14: 2284-2295
- PubMed: 16131657
- DOI: https://doi.org/10.1110/ps.051582605
- Primary Citation of Related Structures:
1YOB - PubMed Abstract:
Flavodoxin II from Azotobacter vinelandii is a "long-chain" flavodoxin and has one of the lowest E1 midpoint potentials found within the flavodoxin family. To better understand the relationship between structural features and redox potentials, the oxidized form of the C69A mutant of this flavodoxin was crystallized and its three-dimensional structure determined to a resolution of 2.25 A by molecular replacement. Its overall fold is similar to that of other flavodoxins, with a central five-stranded parallel beta-sheet flanked on either side by alpha-helices. An eight-residue insertion, compared with other long-chain flavodoxins, forms a short 3(10) helix preceding the start of the alpha3 helix. The flavin mononucleotide (FMN) cofactor is flanked by a leucine on its re face instead of the more conserved tryptophan, resulting in a more solvent-accessible FMN binding site and stabilization of the hydroquinone (hq) state. In particular the absence of a hydrogen bond to the N5 atom of the oxidized FMN was identified, which destabilizes the ox form, as well as an exceptionally large patch of acidic residues in the vicinity of the FMN N1 atom, which destabilizes the hq form. It is also argued that the presence of a Gly at position 58 in the sequence stabilizes the semiquinone (sq) form, as a result, raising the E2 value in particular.
- Leiden Institute of Chemistry, P.O. Box 9502, 2300 RA Leiden, The Netherlands.
Organizational Affiliation: