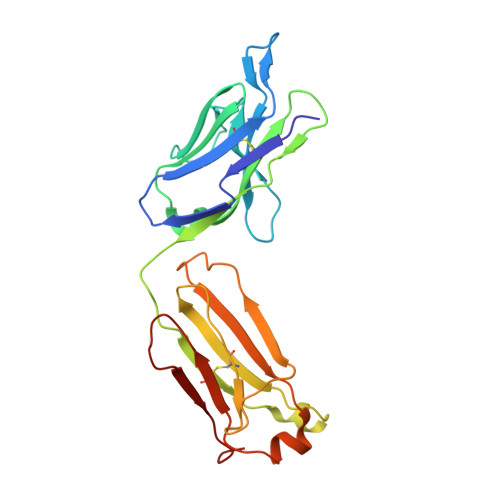
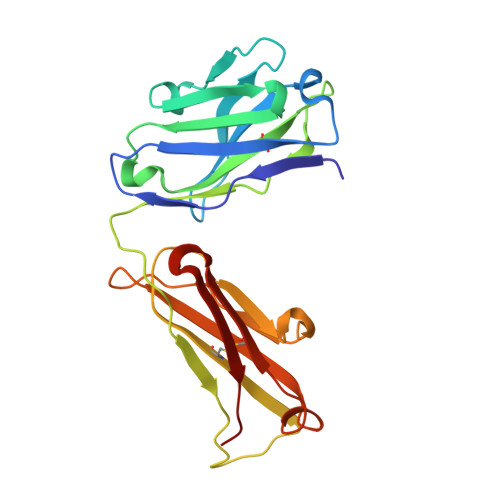
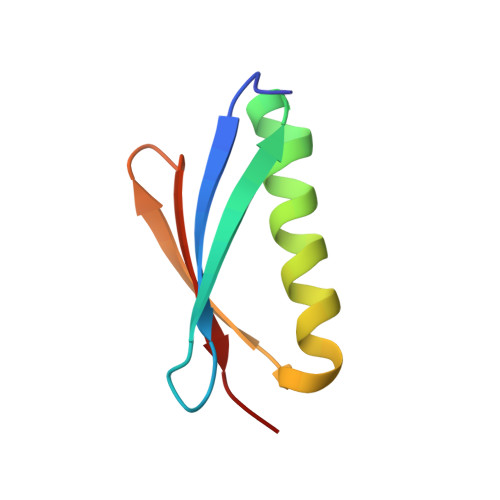
Comparison of the crystallization and crystal packing of two Fab single-site mutant protein L complexes.
Granata, V., Housden, N.G., Harrison, S., Jolivet-Reynaud, C., Gore, M.G., Stura, E.A.(2005) Acta Crystallogr D Biol Crystallogr 61: 750-754
- PubMed: 15930633
- DOI: https://doi.org/10.1107/S0907444905007110
- Primary Citation Related Structures:
1YMH - PubMed Abstract:
Protein L from Peptostreptococcus magnus (PpL) is a multidomain protein composed of four or five immunoglobulin-binding domains that target the kappa light chain of a large repertoire of human and murine antibodies. Thus, a single domain of this protein can be used to aid the crystallization of Fab, free or complexed to their antigen when it is not possible to obtain crystals without it. Each wild-type PpL domain has two light-chain binding sites that target the same region of the light chain and can thus bring together two Fab-antigen complexes within the crystal lattice. In this context the small PpL domain is sandwiched between two Fab and cannot participate in crystal contacts, thus mutants are unlikely to increase the chances of crystallizing a particular complex. However, it is possible to design mutants that can bind at only one site by making use of the crystal structures obtained so far. Such mutants will have a free surface that can participate in crystal contacts and that can be modified to improve its crystal contact-forming properties. Here, a comparison of two single-site mutants that differ at three different positions is reported. In both mutants two different tryptophan residues participate in crystal-packing interactions, suggesting that this residue may be particularly interesting for enhancing crystal-contact formation.
- Università Federico II, Dipartimento di Chimica Biologica, Via Mezzocannone, Napoli, Italy.
Organizational Affiliation: