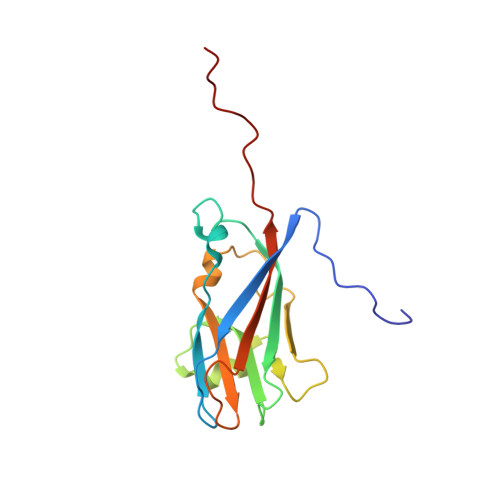
Crystallographic structure of the T=1 particle of brome mosaic virus.
Larson, S.B., Lucas, R.W., McPherson, A.(2005) J Mol Biology 346: 815-831
- PubMed: 15713465 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.12.015
- Primary Citation Related Structures:
1YC6 - PubMed Abstract:
T=1 icosahedral particles of amino terminally truncated brome mosaic virus (BMV) protein were created by treatment of the wild-type T=3 virus with 1M CaCl2 and crystallized from sodium malonate. Diffraction data were collected from frozen crystals to beyond 2.9 A resolution and the structure determined by molecular replacement and phase extension. The particles are composed of pentameric capsomeres from the wild-type virions which have reoriented with respect to the original particle pentameric axes by rotations of 37 degrees , and formed tenuous interactions with one another, principally through conformationally altered C-terminal polypeptides. Otherwise, the pentamers are virtually superimposable upon those of the original T=3 BMV particles. The T=1 particles, in the crystals, are not perfect icosahedra, but deviate slightly from exact symmetry, possibly due to packing interactions. This suggests that the T=1 particles are deformable, which is consistent with the loose arrangement of pentamers and latticework of holes that penetrate the surface. Atomic force microscopy showed that the T=3 to T=1 transition could occur by shedding of hexameric capsomeres and restructuring of remaining pentamers accompanied by direct condensation. Knowledge of the structures of the BMV wild-type and T=1 particles now permit us to propose a tentative model for that process. A comparison of the BMV T=1 particles was made with the reassembled T=1 particles produced from the coat protein of trypsin treated alfalfa mosaic virus (AlMV), another bromovirus. There is little resemblance between the two particles. The BMV particle, with a maximum diameter of 195 A, is made from distinctive pentameric capsomeres with large holes along the 3-fold axis, while the AlMV particle, of approximate maximum diameter 220 A, has subunits closely packed around the 3-fold axis, large holes along the 5-fold axis, and few contacts within pentamers. In both particles crucial linkages are made about icosahedral dyads.
- Department of Molecular Biology and Biochemistry, University of California, Irvine, CA 92697-3900, USA.
Organizational Affiliation: