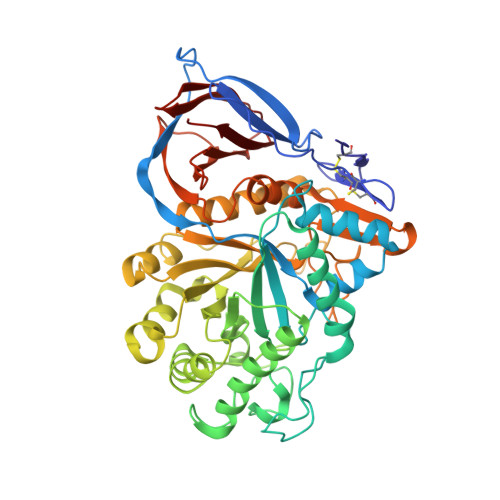
X-ray structure of human acid-beta-glucosidase covalently bound to conduritol-B-epoxide. Implications for Gaucher disease.
Premkumar, L., Sawkar, A.R., Boldin-Adamsky, S., Toker, L., Silman, I., Kelly, J.W., Futerman, A.H., Sussman, J.L.(2005) J Biological Chem 280: 23815-23819
- PubMed: 15817452
- DOI: https://doi.org/10.1074/jbc.M502799200
- Primary Citation of Related Structures:
1Y7V - PubMed Abstract:
Gaucher disease is an inherited metabolic disorder caused by mutations in the lysosomal enzyme acid-beta-glucosidase (GlcCerase). We recently determined the x-ray structure of GlcCerase to 2.0 A resolution (Dvir, H., Harel, M., McCarthy, A. A., Toker, L., Silman, I., Futerman, A. H., and Sussman, J. L. (2003) EMBO Rep.4, 704-709) and have now solved the structure of Glc-Cerase conjugated with an irreversible inhibitor, conduritol-B-epoxide (CBE). The crystal structure reveals that binding of CBE to the active site does not induce a global conformational change in GlcCerase and confirms that Glu340 is the catalytic nucleophile. However, only one of two alternative conformations of a pair of flexible loops (residues 345-349 and 394-399) located at the entrance to the active site in native GlcCerase is observed in the GlcCerase-CBE structure, a conformation in which the active site is accessible to CBE. Analysis of the dynamics of these two alternative conformations suggests that the two loops act as a lid at the entrance to the active site. This possibility is supported by a cluster of mutations in loop 394-399 that cause Gaucher disease by reducing catalytic activity. Moreover, in silico mutational analysis demonstrates that all these mutations stabilize the conformation that limits access to the active site, thus providing a mechanistic explanation of how mutations in this loop result in Gaucher disease.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: