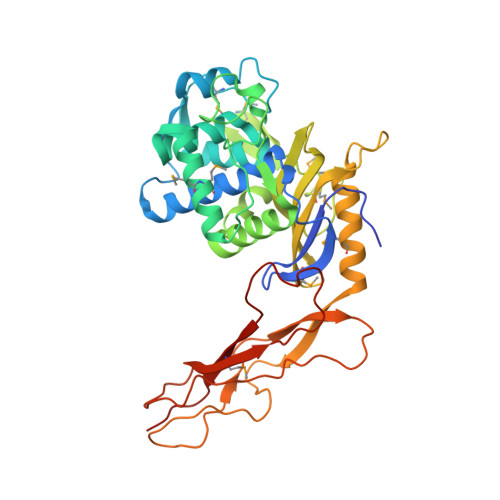
Crystal structure of a peptidoglycan synthesis regulatory factor (PBP3) from Streptococcus pneumoniae
Morlot, C., Pernot, L., Le Gouellec, A., Di Guilmi, A.M., Vernet, T., Dideberg, O., Dessen, A.(2005) J Biological Chem 280: 15984-15991
- PubMed: 15596446
- DOI: https://doi.org/10.1074/jbc.M408446200
- Primary Citation Related Structures:
1XP4 - PubMed Abstract:
Penicillin-binding proteins (PBPs) are membrane-associated enzymes which perform critical functions in the bacterial cell division process. The single d-Ala,d-Ala (d,d)-carboxypeptidase in Streptococcus pneumoniae, PBP3, has been shown to play a key role in control of availability of the peptidoglycal substrate during cell growth. Here, we have biochemically characterized and solved the crystal structure of a soluble form of PBP3 to 2.8 A resolution. PBP3 folds into an NH(2)-terminal, d,d-carboxypeptidase-like domain, and a COOH-terminal, elongated beta-rich region. The carboxypeptidase domain harbors the classic signature of the penicilloyl serine transferase superfamily, in that it contains a central, five-stranded antiparallel beta-sheet surrounded by alpha-helices. As in other carboxypeptidases, which are present in species whose peptidoglycan stem peptide has a lysine residue at the third position, PBP3 has a 14-residue insertion at the level of its omega loop, a feature that distinguishes it from carboxypeptidases from bacteria whose peptidoglycan harbors a diaminopimelate moiety at this position. PBP3 performs substrate acylation in a highly efficient manner (k(cat)/K(m) = 50,500 M(-1) x s(-1)), an event that may be linked to role in control of pneumococcal peptidoglycan reticulation. A model that places PBP3 poised vertically on the bacterial membrane suggests that its COOH-terminal region could act as a pedestal, placing the active site in proximity to the peptidoglycan and allowing the protein to "skid" on the surface of the membrane, trimming pentapeptides during the cell growth and division processes.
- Laboratoire de Cristallographie Macromoléculaire and Laboratoire d'Ingénierie des Macromolécules, Institut de Biologie Structurale Jean-Pierre Ebel (CNRS/CEA/UJF), 41 rue Jules Horowitz, Grenoble 38027, France.
Organizational Affiliation: