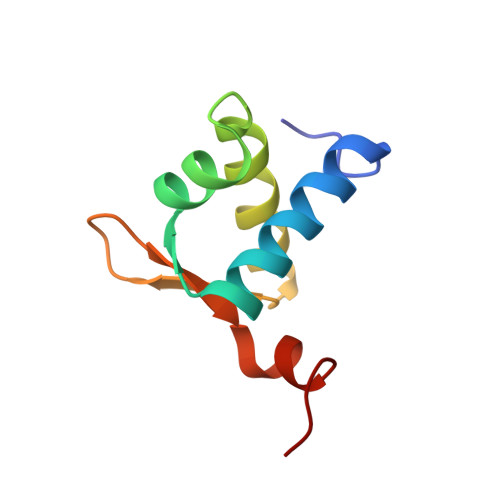
The Crystal Structure of the Z[beta] Domain of the RNA-editing Enzyme ADAR1 Reveals Distinct Conserved Surfaces Among Z-domains.
Athanasiadis, A., Placido, D., Maas, S., Brown II, B.A., Lowenhaupt, K., Rich, A.(2005) J Mol Biology 351: 496-507
- PubMed: 16023667 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.06.028
- Primary Citation Related Structures:
1XMK - PubMed Abstract:
The Zalpha domains represent a growing subfamily of the winged helix-turn-helix (HTH) domain family whose members share a remarkable ability to bind specifically to Z-DNA and/or Z-RNA. They have been found exclusively in proteins involved in interferon response and, while their importance in determining pox viral pathogenicity has been demonstrated, their actual target and biological role remain obscure. Cellular proteins containing Zalpha domains bear a second homologous domain termed Zbeta, which appears to lack the ability to bind left-handed nucleic acids. Here, we present the crystal structure of the Zbeta domain from the human double-stranded RNA adenosine deaminase ADAR1 at 0.97 A, determined by single isomorphous replacement including anomalous scattering. Zbeta maintains a winged-HTH fold with the addition of a C-terminal helix. Mapping of the Zbeta conservation profile on the Zbeta surface reveals a new conserved surface formed partly by the terminal helix 4, involved in metal binding and dimerization and absent from Zalpha domains. Our results show how two domains similar in fold may have evolved into different functional entities even in the context of the same protein.
- Department of Biology, Massachusetts Institute of Technology, 77 Massachusetts Avenue, Cambridge, MA 02139, USA. alekosathanasiadis@hotmail.com
Organizational Affiliation: