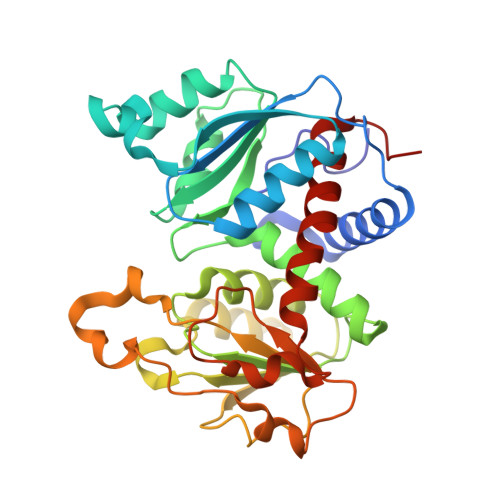
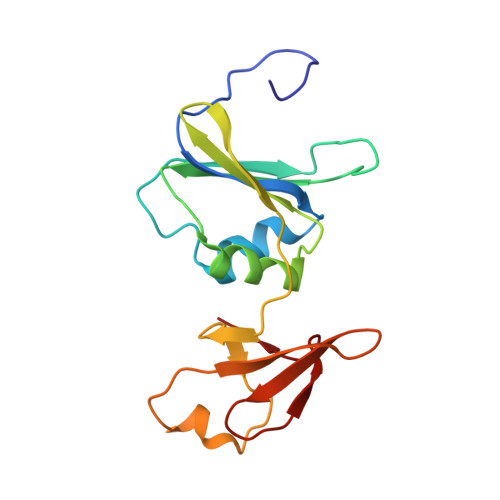
A Single Amino Acid Substitution in the Active Site of Escherichia coli Aspartate Transcarbamoylase Prevents the Allosteric Transition.
Stieglitz, K.A., Pastra-Landis, S.C., Xia, J., Tsuruta, H., Kantrowitz, E.R.(2005) J Mol Biology 349: 413-423
- PubMed: 15890205 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2005.03.073
- Primary Citation Related Structures:
1XJW - PubMed Abstract:
Modeling of the tetrahedral intermediate within the active site of Escherichia coli aspartate transcarbamoylase revealed a specific interaction with the side-chain of Gln137, an interaction not previously observed in the structure of the X-ray enzyme in the presence of N-phosphonacetyl-L-aspartate (PALA). Previous site-specific mutagenesis experiments showed that when Gln137 was replaced by alanine, the resulting mutant enzyme (Q137A) exhibited approximately 50-fold less activity than the wild-type enzyme, exhibited no homotropic cooperativity, and the binding of both carbamoyl phosphate and aspartate were extremely compromised. To elucidate the structural alterations in the mutant enzyme that might lead to such pronounced changes in kinetic and binding properties, the Q137A enzyme was studied by time-resolved, small-angle X-ray scattering and its structure was determined in the presence of PALA to 2.7 angstroms resolution. Time-resolved, small-angle X-ray scattering established that the natural substrates, carbamoyl phosphate and L-aspartate, do not induce in the Q137A enzyme the same conformational changes as observed for the wild-type enzyme, although the scattering pattern of the Q137A and wild-type enzymes in the presence of PALA were identical. The overall structure of the Q137A enzyme is similar to that of the R-state structure of wild-type enzyme with PALA bound. However, there are differences in the manner by which the Q137A enzyme coordinates PALA, especially in the side-chain positions of Arg105 and His134. The replacement of Gln137 by Ala also has a dramatic effect on the electrostatics of the active site. These data taken together suggest that the side-chain of Gln137 in the wild-type enzyme is required for the binding of carbamoyl phosphate in the proper orientation so as to induce conformational changes required for the creation of the high-affinity aspartate-binding site. The inability of carbamoyl phosphate to create the high-affinity binding site in the Q137A enzyme results in an enzyme locked in the low-activity low-affinity T state. These results emphasize the absolute requirement of the binding of carbamoyl phosphate for the creation of the high-affinity aspartate-binding site and for inducing the homotropic cooperativity in aspartate transcarbamoylase.
- Department of Chemistry, Merkert Chemistry Center, Boston College, Chestnut Hill, MA 02467, USA.
Organizational Affiliation: