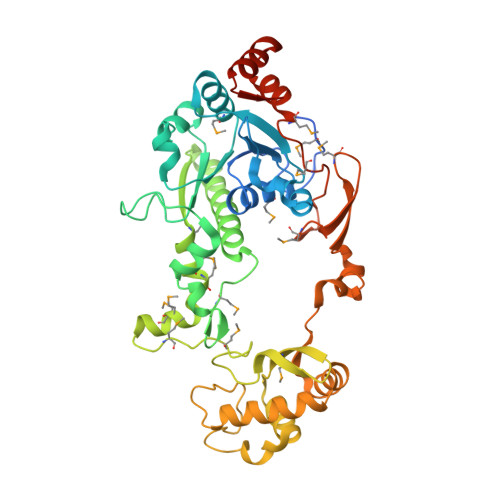
Crystal Structure of CaiB, a Type-III CoA Transferase in Carnitine Metabolism
Stenmark, P., Gurmu, D., Nordlund, P.(2004) Biochemistry 43: 13996-14003
- PubMed: 15518548 Search on PubMed
- DOI: https://doi.org/10.1021/bi048481c
- Primary Citation Related Structures:
1XA3, 1XA4 - PubMed Abstract:
Carnitine is an important molecule in human metabolism, mainly because of its role in the transport of long-chain fatty acids across the inner mitochondrial membrane. Escherichia coli uses carnitine as a terminal electron acceptor during anaerobic metabolism. Bacteria present in our large intestine break down carnitine that is not absorbed in the small intestine. One part of this catabolic pathway is reversible and can be utilized for bioproduction of large amounts of stereochemically pure L-carnitine, which is used medically for the treatment of a variety of human diseases. Here, we present the crystal structure of the E. coli protein CaiB, which is a member of the recently identified type-III coenzyme A (CoA) transferase family and catalyzes the transfer of the CoA moiety between gamma-butyrobetaine-CoA and carnitine forming carnityl-CoA and gamma-butyrobetaine. This is the first protein from the carnitine metabolic pathway to be structurally characterized. The structure of CaiB reveals a spectacular fold where two monomers are interlaced to form an interlocked dimer. A molecule of the crystallization buffer bis-(2-hydroxyethyl)imino-tris(hydroxymethyl)methane (bis-tris) is bound in a large pocket located primarily in the small domain, and we propose that this pocket constitutes the binding site for both substrate moieties participating in the CaiB transfer reaction. The binding of CoA to CaiB induces a domain movement that closes the active site of the protein. This is the first observation of a domain movement in the type-III CoA transferase family and can play an important role in coupling substrate binding to initiation of the catalytic reaction.
- Department of Biochemistry and Biophysics, Stockholm University, Roslagstullsbacken 15, Albanova University Center, SE-10691 Stockholm, Sweden.
Organizational Affiliation: