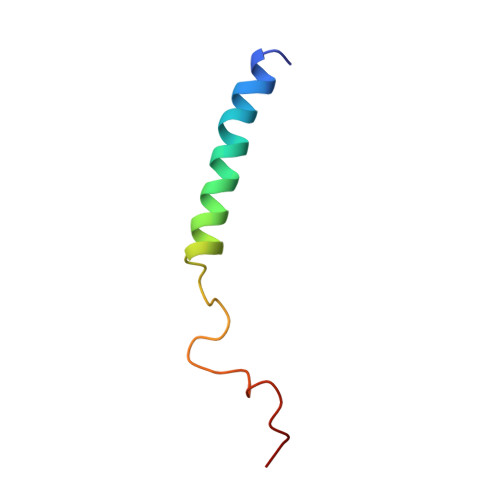
The C-terminal domain of the HIV-1 regulatory protein Vpr adopts an antiparallel dimeric structure in solution via its leucine-zipper-like domain
Bourbigot, S., Beltz, H., Denis, J., Morellet, N., Roques, B.P., Mely, Y., Bouaziz, S.(2005) Biochem J 387: 333-341
- PubMed: 15571493
- DOI: https://doi.org/10.1042/BJ20041759
- Primary Citation of Related Structures:
1X9V - PubMed Abstract:
HIV-1 Vpr is a highly conserved accessory protein that is involved in many functions of the virus life cycle. Vpr facilitates the entry of the HIV pre-integration complex through the nuclear pore, induces G2 cell cycle arrest, regulates cell apoptosis, increases transcription from the long terminal repeat and enhances viral replication. Vpr contains a Leu/Ile-rich domain (amino acids 60-81) in its C-terminal part, which is critical for dimerization. The sequence comprising residues 52-96 is implicated in properties of the protein such as DNA interaction and apoptosis via interaction with the adenine nucleotide translocator. To understand the specific interactions of Vpr-(52-96), the ability of this peptide to dimerize via a leucine-zipper mechanism has been investigated, by NMR and fluorescence spectroscopy. In contrast with results from a study performed in the presence of trifluoroethanol, our results, obtained in 30% (v/v) [2H]acetonitrile, show that Vpr-(52-96) in solution still forms an a-helix spanning residues 53-75, but dimerizes in an antiparallel orientation, through hydrophobic interactions between leucine and isoleucine residues and stacking between His71 and Trp54. Moreover, to demonstrate the physiological relevance of the dimer structure, fluorescence spectroscopy experiments have been performed in a Mes buffer, which confirmed the formation of the dimer in aqueous solution and highlighted the spatial proximity between Trp54 and His71. Surprisingly, the leucine-zipper structure shown in the present work for Vpr-(52-96) mimics the structure of full-length Vpr-(1-96), and this could explain why some of the properties of Vpr-(52-96) and Vpr-(1-96) are identical, while some are even enhanced for Vpr-(52-96), particularly in the case of DNA transfection experiments.
- Département de Pharmacologie Chimique & Génétique, INSERM U640-CNRS UMR 8151, UFR des Sciences Pharmaceutiques et Biologiques, 4 avenue de l'Observatoire, 75270 Paris Cedex 06, France.
Organizational Affiliation: