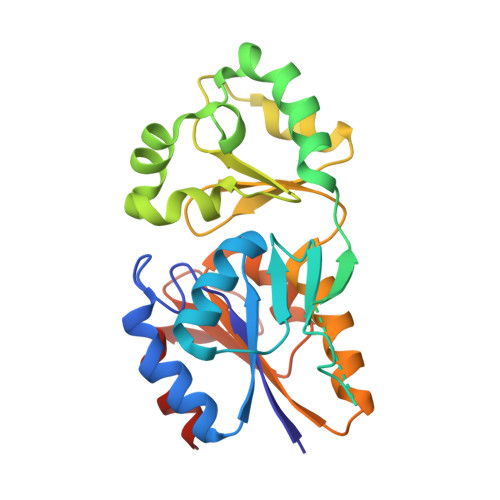
Structure of mannosyl-3-phosphoglycerate phosphatase from Pyrococcus horikoshii.
Kawamura, T., Watanabe, N., Tanaka, I.(2008) Acta Crystallogr D Biol Crystallogr 64: 1267-1276
- PubMed: 19018103
- DOI: https://doi.org/10.1107/S0907444908033817
- Primary Citation of Related Structures:
1WZC, 2ZOS - PubMed Abstract:
Mannosyl-3-phosphoglycerate phosphatase (MPGP) catalyzes the dephosphorylation of alpha-mannosyl-3-phosphoglycerate (MPG) to produce alpha-mannosylglycerate (MG). In the hyperthermophile Pyrococcus horikoshii, MPGP plays a role in a series of enzyme reactions that are involved in the MG-biosynthesis pathway, which is important for maintaining life under conditions of high salt concentration. Crystal structures of P. horikoshii MPGP (PhoMPGP) in the holo form and in the apo form lacking the magnesium ion were determined by the multiple-wavelength anomalous diffraction method using SeMet-substituted PhoMPGP. PhoMPGP consists of two domains: a core domain that is conserved in the haloacid dehalogenase superfamily and a cap domain that is specific to the C2B cap subclass of the superfamily. Apo-form crystals contain two PhoMPGP molecules: one in the open conformation and the other in the closed conformation. In holo-form crystals both of the two molecules are in the closed conformation with phosphate and magnesium ions. PhoMPGP has a specific hairpin loop that is bent towards the active site in the closed conformation of both the apo and holo forms. PhoMPGP has a cavity between the two domains which is considered to be the substrate-binding site as a phosphate ion is located in the cavity, mimicking the binding manner of the phosphate group of MPG. The cavity is sequestered in the closed conformation such that a conformational change is indispensable for the release of products. A salt bridge from the general acid/base Asp10 to Arg170 is observed in the holo-form PhoMPGP which is not present in the open form. The importance of the conformational change in the activity of PhoMPGP is discussed.
- Division of Biological Sciences, Graduate School of Science, Hokkaido University, Sapporo 060-0810, Japan.
Organizational Affiliation: