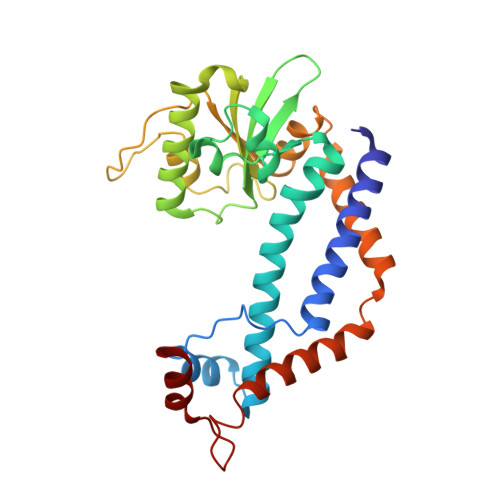
Crystal structures of type II restriction endonuclease EcoO109I and its complex with cognate DNA
Hashimoto, H., Shimizu, T., Imasaki, T., Kato, M., Shichijo, N., Kita, K., Sato, M.(2005) J Biological Chem 280: 5605-5610
- PubMed: 15590682 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M411684200
- Primary Citation Related Structures:
1WTD, 1WTE - PubMed Abstract:
EcoO109I is a type II restriction endonuclease that recognizes the DNA sequence of RGGNCCY. Here we describe the crystal structures of EcoO109I and its complex with DNA. A comparison of the two structures shows that the catalytic domain moves drastically to capture the DNA. One metal ion and two water molecules are observed near the active site of the DNA complex. The metal ion is a Lewis acid that stabilizes the pentavalent phosphorus atom in the transition state. One water molecule, activated by Lys-126, attacks the phosphorus atom in an S(N)2 mechanism, whereas the other water interacts with the 3'-leaving oxygen to donate a proton to the oxygen. EcoO109I is similar to EcoRI family enzymes in terms of its DNA cleavage pattern and folding topology of the common motif in the catalytic domain, but it differs in the manner of DNA recognition. Our findings propose a novel classification of the type II restriction endonucleases and lead to the suggestion that EcoO109I represents a new subclass of the EcoRI family.
- Graduate School of Integrated Science, Yokohama City University, 1-7-29 Suehiro-cho, Tsurumi-ku, Yokohama 230-0045, Japan.
Organizational Affiliation: