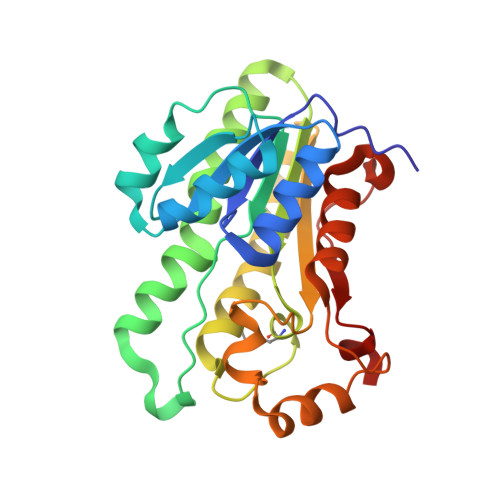
Structure of the tetrameric form of human L-Xylulose reductase: Probing the inhibitor-binding site with molecular modeling and site-directed mutagenesis
El-Kabbani, O., Carbone, V., Darmanin, C., Ishikura, S., Hara, A.(2005) Proteins 60: 424-432
- PubMed: 15906319 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20487
- Primary Citation Related Structures:
1WNT - PubMed Abstract:
L-Xylulose reductase (XR) is a member of the short-chain dehydrogenase/reductase (SDR) superfamily. In this study we report the structure of the biological tetramer of human XR in complex with NADP(+) and a competitive inhibitor solved at 2.3 A resolution. A single subunit of human XR is formed by a centrally positioned, seven-stranded, parallel beta-sheet surrounded on either side by two arrays of three alpha-helices. Two helices located away from the main body of the protein form the variable substrate-binding cleft, while the dinucleotide coenzyme-binding motif is formed by a classical Rossmann fold. The tetrameric structure of XR, which is held together via salt bridges formed by the guanidino group of Arg203 from one monomer and the carboxylate group of the C-terminal residue Cys244 from the neighboring monomer, explains the ability of human XR to prevent the cold inactivation seen in the rodent forms of the enzyme. The orientations of Arg203 and Cys244 are maintained by a network of hydrogen bonds and main-chain interactions of Gln137, Glu238, Phe241, and Trp242. These interactions are similar to those defining the quaternary structure of the closely related carbonyl reductase from mouse lung. Molecular modeling and site-directed mutagenesis identified the active site residues His146 and Trp191 as forming essential contacts with inhibitors of XR. These results could provide a structural basis in the design of potent and specific inhibitors for human XR.
- Department of Medicinal Chemistry, Victorian College of Pharmacy, Monash University, Parkville, Victoria, Australia. ossama.el-kabbani@vcp.monash.edu.au
Organizational Affiliation: