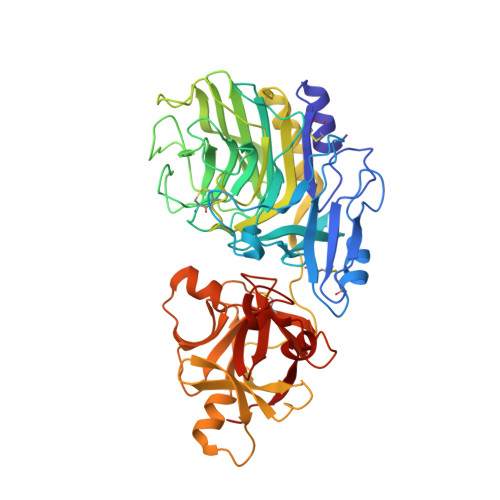
Crystal structure of a family 54 alpha-L-arabinofuranosidase reveals a novel carbohydrate-binding module that can bind arabinose
Miyanaga, A., Koseki, T., Matsuzawa, H., Wakagi, T., Shoun, H., Fushinobu, S.(2004) J Biological Chem 279: 44907-44914
- PubMed: 15292273
- DOI: https://doi.org/10.1074/jbc.M405390200
- Primary Citation of Related Structures:
1WD3, 1WD4 - PubMed Abstract:
As the first known structures of a glycoside hydrolase family 54 (GH54) enzyme, we determined the crystal structures of free and arabinose-complex forms of Aspergillus kawachii IFO4308 alpha-l-arabinofuranosidase (AkAbfB). AkAbfB comprises two domains: a catalytic domain and an arabinose-binding domain (ABD). The catalytic domain has a beta-sandwich fold similar to those of clan-B glycoside hydrolases. ABD has a beta-trefoil fold similar to that of carbohydrate-binding module (CBM) family 13. However, ABD shows a number of characteristics distinctive from those of CBM family 13, suggesting that it could be classified into a new CBM family. In the arabinose-complex structure, one of three arabinofuranose molecules is bound to the catalytic domain through many interactions. Interestingly, a disulfide bond formed between two adjacent cysteine residues recognized the arabinofuranose molecule in the active site. From the location of this arabinofuranose and the results of a mutational study, the nucleophile and acid/base residues were determined to be Glu(221) and Asp(297), respectively. The other two arabinofuranose molecules are bound to ABD. The O-1 atoms of the two arabinofuranose molecules bound at ABD are both pointed toward the solvent, indicating that these sites can both accommodate an arabinofuranose side-chain moiety linked to decorated arabinoxylans.
- Department of Biotechnology, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: