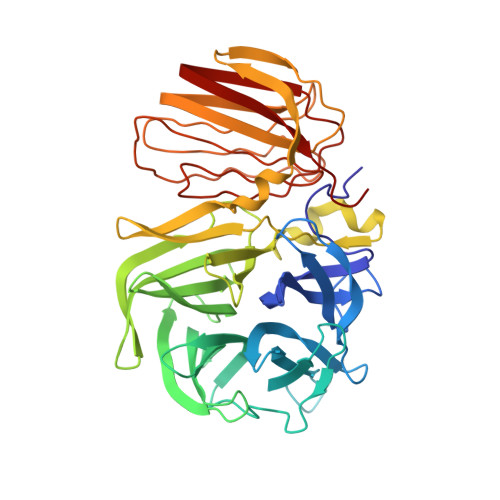
Crystal Structure of Inactivated Thermotoga Maritima Invertase in Complex with the Trisaccharide Substrate Raffinose.
Alberto, F., Jordi, E., Henrissat, B., Czjzek, M.(2006) Biochem J 395: 457
- PubMed: 16411890
- DOI: https://doi.org/10.1042/BJ20051936
- Primary Citation of Related Structures:
1W2T - PubMed Abstract:
Thermotoga maritima invertase (beta-fructosidase), a member of the glycoside hydrolase family GH-32, readily releases beta-D-fructose from sucrose, raffinose and fructan polymers such as inulin. These carbohydrates represent major carbon and energy sources for prokaryotes and eukaryotes. The invertase cleaves beta-fructopyranosidic linkages by a double-displacement mechanism, which involves a nucleophilic aspartate and a catalytic glutamic acid acting as a general acid/base. The three-dimensional structure of invertase shows a bimodular enzyme with a five bladed beta-propeller catalytic domain linked to a beta-sandwich of unknown function. In the present study we report the crystal structure of the inactivated invertase in interaction with the natural substrate molecule alpha-D-galactopyranosyl-(1,6)-alpha-D-glucopyranosyl-beta-D-fructofuranoside (raffinose) at 1.87 A (1 A=0.1 nm) resolution. The structural analysis of the complex reveals the presence of three binding-subsites, which explains why T. maritima invertase exhibits a higher affinity for raffinose than sucrose, but a lower catalytic efficiency with raffinose as substrate than with sucrose.
- Architecture et Fonction des Macromolécules Biologiques, UMR6098, CNRS, Universités Aix-Marseille I & II, Case 932, 163 Avenue de Luminy, 13288 Marseille Cedex 9, France.
Organizational Affiliation: