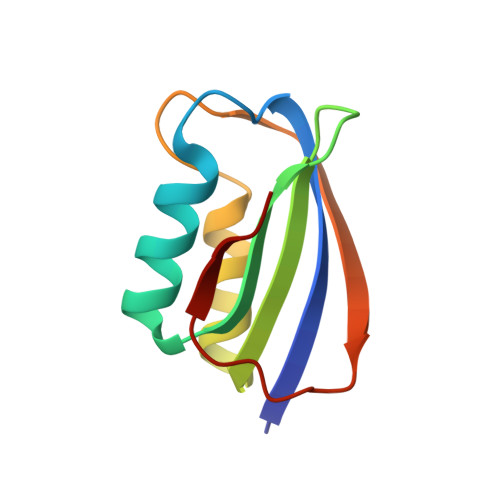
Crystal Structure of a Hyperthermophilic Archaeal Acylphosphatase from Pyrococcus Horikoshii-Structural Insights Into Enzymatic Catalysis, Thermostability, and Dimerization
Cheung, Y.Y., Lam, S.Y., Chu, W.K., Allen, M.D., Bycroft, M., Wong, K.B.(2005) Biochemistry 44: 4601
- PubMed: 15779887
- DOI: https://doi.org/10.1021/bi047832k
- Primary Citation of Related Structures:
1W2I - PubMed Abstract:
Acylphosphatases catalyze the hydrolysis of the carboxyl-phosphate bond in acyl phosphates. Although acylphosphatase-like sequences are found in all three domains of life, no structure of acylphosphatase has been reported for bacteria and archaea so far. Here, we report the characterization of enzymatic activities and crystal structure of an archaeal acylphosphatase. A putative acylphosphatase gene (PhAcP) was cloned from the genomic DNA of Pyrococcus horikoshii and was expressed in Escherichia coli. Enzymatic parameters of the recombinant PhAcP were measured using benzoyl phosphate as the substrate. Our data suggest that, while PhAcP is less efficient than other mammalian homologues at 25 degrees C, the thermophilic enzyme is fully active at the optimal growth temperature (98 degrees C) of P. horikoshii. PhAcP is extremely stable; its apparent melting temperature was 111.5 degrees C and free energy of unfolding at 25 degrees C was 54 kJ mol(-)(1). The 1.5 A crystal structure of PhAcP adopts an alpha/beta sandwich fold that is common to other acylphosphatases. PhAcP forms a dimer in the crystal structure via antiparallel association of strand 4. Structural comparison to mesophilic acylphosphatases reveals significant differences in the conformation of the L5 loop connecting strands 4 and 5. The extreme thermostability of PhAcP can be attributed to an extensive ion-pair network consisting of 13 charge residues on the beta sheet of the protein. The reduced catalytic efficiency of PhAcP at 25 degrees C may be due to a less flexible active-site residue, Arg20, which forms a salt bridge to the C-terminal carboxyl group. New insights into catalysis were gained by docking acetyl phosphate to the active site of PhAcP.
- Department of Biochemistry, The Chinese University of Hong Kong, Hong Kong, China.
Organizational Affiliation: