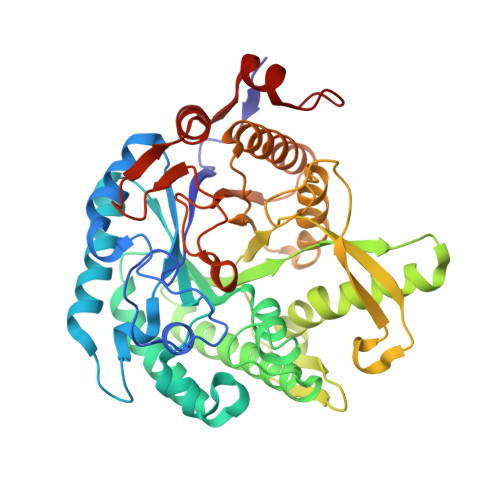
X-ray structure of a membrane-bound beta-glycosidase from the hyperthermophilic archaeon Pyrococcus horikoshii
Akiba, T., Nishio, M., Matsui, I., Harata, K.(2004) Proteins 57: 422-431
- PubMed: 15340929 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20203
- Primary Citation Related Structures:
1VFF - PubMed Abstract:
The beta-glycosidase of the hyperthermophilic Archaeon Pyrococcus horikoshii is a membrane-bound enzyme with the preferred substrate of alkyl-beta-glycosides. In this study, the unusual structural features that confer the extreme thermostability and substrate preferences of this enzyme were investigated by X-ray crystallography and docking simulation. The enzyme was crystallized in the presence of a neutral surfactant, and the crystal structure was solved by the molecular replacement method and refined at 2.5 A. The main-chain fold of the enzyme belongs to the (betaalpha)8 barrel structure common to the Family 1 glycosyl hydrolases. The active site is located at the center of the C-termini of the barrel beta-strands. The deep pocket of the active site accepts one sugar unit, and a hydrophobic channel extending radially from there binds the nonsugar moiety of the substrate. The docking simulation for oligosaccharides and alkylglucosides indicated that alkylglucosides with a long aliphatic chain are easily accommodated in the hydrophobic channel. This sparingly soluble enzyme has a cluster of hydrophobic residues on its surface, situated at the distal end of the active site channel and surrounded by a large patch of positively charged residues. We propose that this hydrophobic region can be inserted into the membrane while the surrounding positively charged residues make favorable contacts with phosphate groups on the inner surface of the membrane. The enzyme could thus adhere to the membrane in the proximity of its glycolipid substrate.
- Biological Information Research Center, National Institute of Advanced Industrial Science and Technology, Tsukuba, Japan.
Organizational Affiliation: