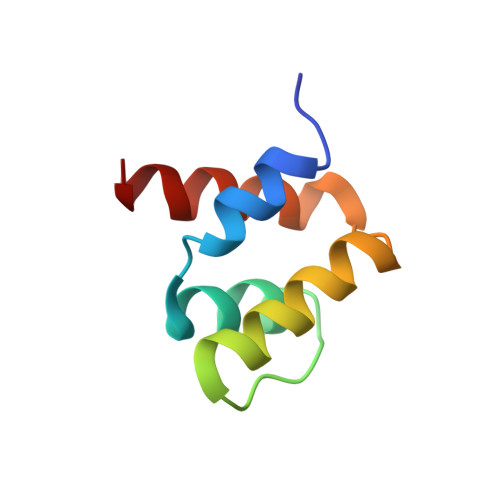
NMR structure of the N-terminal domain of SUMO ligase PIAS1 and its interaction with tumor suppressor p53 and A/T-rich DNA oligomers
Okubo, S., Hara, F., Tsuchida, Y., Shimotakahara, S., Suzuki, S., Hatanaka, H., Yokoyama, S., Tanaka, H., Yasuda, H., Shindo, H.(2004) J Biological Chem 279: 31455-31461
- PubMed: 15133049
- DOI: https://doi.org/10.1074/jbc.M403561200
- Primary Citation Related Structures:
1V66 - PubMed Abstract:
A member of the PIAS (protein inhibitor of activated STAT) family of proteins, PIAS1, have been reported to serve as an E3-type SUMO ligase for tumor suppressor p53 and its own. It also was proposed that the N-terminal domain of PIAS1 interacts with DNA as well as p53. Extensive biochemical studies have been devoted recently to understand sumoylations and its biological implications, whereas the structural aspects of the PIAS family and the mechanism of its interactions with various factors are less well known to date. In this study, the three-dimensional structure of the N-terminal domain (residues 1-65) of SUMO ligase PIAS1 was determined by NMR spectroscopy. The structure revealed a unique four-helix bundle with a topology of an up-down-extended loop-down-up, a part of which the helix-extended loop-helix represented the SAP (SAF-A/B, Acinus, and PIAS) motif. Thus, this N-terminal domain may be referred to as a four-helix SAP domain. The glutathione S-transferase pull-down assay demonstrated that this domain possesses a binding ability to tumor suppressor p53, a target protein for sumoylation by PIAS1, whereas gel mobility assays showed that it has a strong affinity toward A/T-rich DNA. An NMR analysis of the four-helix SAP domain complexed with the 16-bp-long DNA demonstrated that one end of the four-helix bundle is the binding site and may fit into the minor groove of DNA. The three-dimensional structure and its binding duality are discussed in conjunction with the biological functions of PIAS1 as a SUMO ligase.
- School of Pharmacy, Tokyo University of Pharmacy & Life Science, Horinouchi, Hachioji, Tokyo 192-0392, Japan.
Organizational Affiliation: