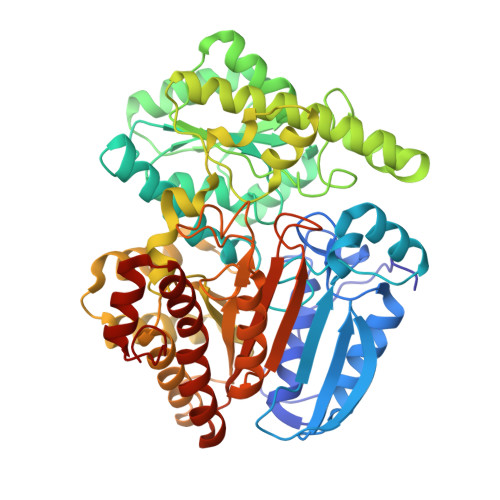
Structure and Action of Urocanase
Kessler, D., Retey, J., Schulz, G.E.(2004) J Mol Biology 342: 183
- PubMed: 15313616 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.07.028
- Primary Citation Related Structures:
1UWK, 1UWL, 1W1U - PubMed Abstract:
Urocanase (EC 4.2.1.49) from Pseudomonas putida was crystallized after removing one of the seven free thiol groups. The crystal structure was solved by multiwavelength anomalous diffraction (MAD) using a seleno-methionine derivative and then refined at 1.14 A resolution. The enzyme is a symmetric homodimer of 2 x 557 amino acid residues with tightly bound NAD+ cofactors. Each subunit consists of a typical NAD-binding domain inserted into a larger core domain that forms the dimer interface. The core domain has a novel chain fold and accommodates the substrate urocanate in a surface depression. The NAD domain sits like a lid on the core domain depression and points with the nicotinamide group to the substrate. Substrate, nicotinamide and five water molecules are completely sequestered in a cavity. Most likely, one of these water molecules hydrates the substrate during catalysis. This cavity has to open for substrate passage, which probably means lifting the NAD domain. The observed atomic arrangement at the active center gives rise to a detailed proposal for the catalytic mechanism that is consistent with published chemical data. As expected, the variability of the residues involved is low, as derived from a family of 58 proteins annotated as urocanases in the data banks. However, one well-embedded member of this family showed a significant deviation at the active center indicating an incorrect annotation.
- Institut für Organische Chemie und Biochemie, Albert-Ludwigs-Universität, Albertstr. 21, 79104 Freiburg im Breisgau, Germany.
Organizational Affiliation: