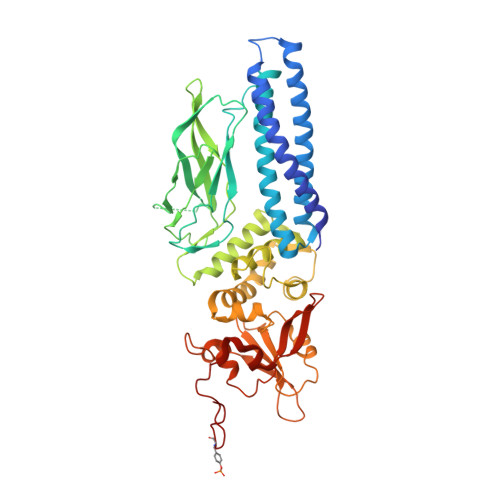
Structure of an Activated Dictyostelium Stat in its DNA-Unbound Form
Soler-Lopez, M., Petosa, C., Fukuzawa, M., Ravelli, R., Williams, J.G., Muller, C.W.(2004) Mol Cell 13: 791
- PubMed: 15053873 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(04)00130-3
- Primary Citation Related Structures:
1UUR, 1UUS - PubMed Abstract:
Dd-STATa is a STAT protein which transcriptionally regulates cellular differentiation in Dictyostelium discoideum, the only non-metazoan known to employ SH2 domain signaling. The 2.7 A crystal structure of a tyrosine phosphorylated Dd-STATa homodimer reveals a four-domain architecture similar to that of mammalian STATs 1 and 3, but with an inverted orientation for the coiled-coil domain. Dimerization is mediated by SH2 domain:phosphopeptide interactions and by a direct interaction between SH2 domains. The unliganded Dd-STATa dimer adopts a fully extended conformation remarkably different from that of the DNA-bound mammalian STATs, implying a large conformational change upon target site recognition. Buried hydrophilic residues predicted to destabilize the coiled-coil domain suggest how hydrophobic residues may become exposed and mediate nuclear export. Functional and evolutionary implications for metazoan STAT proteins are discussed.
- European Molecular Biology Laboratory, Grenoble Outstation, B.P. 181, 38042 Grenoble Cedex 9, France.
Organizational Affiliation: