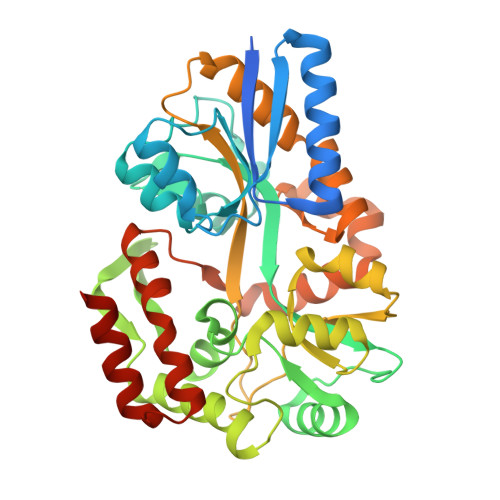
X-Ray Structures of the Maltose-Maltodextrin-Binding Protein of the Thermoacidophilic Bacterium Alicyclobacillus Acidocaldarius Provide Insight Into Acid Stability of Proteins
Schafer, K., Magnusson, U., Scheffel, F., Schiefner, A., Sandgren, M.O.J., Diederichs, K., Welte, W., Hulsmann, A., Schneider, E., Mowbray, S.L.(2004) J Mol Biology 335: 261
- PubMed: 14659755 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.10.042
- Primary Citation Related Structures:
1URD, 1URG, 1URS - PubMed Abstract:
Maltose-binding proteins act as primary receptors in bacterial transport and chemotaxis systems. We report here crystal structures of the thermoacidostable maltose-binding protein from Alicyclobacillus acidocaldarius, and explore its modes of binding to maltose and maltotriose. Further, comparison with the structures of related proteins from Escherichia coli (a mesophile), and two hyperthermophiles (Pyrococcus furiosus and Thermococcus litoralis) allows an investigation of the basis of thermo- and acidostability in this family of proteins.The thermoacidophilic protein has fewer charged residues than the other three structures, which is compensated by an increase in the number of polar residues. Although the content of acidic and basic residues is approximately equal, more basic residues are exposed on its surface whereas most acidic residues are buried in the interior. As a consequence, this protein has a highly positive surface charge. Fewer salt bridges are buried than in the other MBP structures, but the number exposed on its surface does not appear to be unusual. These features appear to be correlated with the acidostability of the A. acidocaldarius protein rather than its thermostability. An analysis of cavities within the proteins shows that the extremophile proteins are more closely packed than the mesophilic one. Proline content is slightly higher in the hyperthermophiles and thermoacidophiles than in mesophiles, and this amino acid is more common at the second position of beta-turns, properties that are also probably related to thermostability. Secondary structural content does not vary greatly in the different structures, and so is not a contributing factor.
- Fachbereich Biologie, Universität Konstanz, Universitätsstrasse 10, 78457, Konstanz, Germany.
Organizational Affiliation: